LEAD ARTICLE
Taylor Medlock-Lanierᵃ ᵇ
ᵃFranklin College of Arts and Sciences, University of Georgia, Athens, GA, USA
ᵇWashington University School of Medicine, St. Louis, MO, USA
CITATION
Medlock-Lanier, Taylor. (2026). Demystifying nervous system regeneration: A comprehensive review. Barton Journal, 1(1), 1–46. https://bartonjournal.org/vol-1-no-1/2026-lead-article-no-001
Abstract
Uncovering mechanisms that govern successful nervous system regeneration will help inform future development of treatments for neurodegenerative disease and brain injury. There are organisms capable of robust nervous system regeneration, but the ability to functionally repair the entire brain is rare. Planarians are freshwater flatworms capable of whole-body regeneration after nearly any injury, including that of their nervous system. Planarians were used as a model to study how successful nervous system regeneration occurs. In this thesis research, six transcription factor-encoding genes were found that are required for the regeneration and maintenance of dopaminergic neurons in the planarian central, peripheral nervous system, and pharyngeal nervous system. Work was done to develop planarians as a model for neurodegenerative disease research through targeted studies and an unbiased screen. The work reviewed here centers around one goal: demystifying nervous system regeneration.
Keywords: regeneration, dopamine, nervous system, planarian, regenerative neurogenesis, neuronal cell fate, specification, transcriptional regulation, differentiation, body patterning, neurodegeneration
Nervous System Development
Development is the process by which a single cell transforms into a complex, multicelled organism. The key stages of development typically include fertilization, cleavage, blastula formation, gastrulation, and organogenesis. Fertilization is when gametes fuse to form a zygote, which will undergo rapid cell division (cleavage) and form the blastula (Gilbert, 2000a, 2000c). During gastrulation, the cells in the blastula will spatially rearrange into three germ layers: the endoderm, mesoderm, and ectoderm (Muhr et al., 2023; Purves et al., 2001b). After gastrulation, the three germ layers will continue to divide and become specialized cell types that form all the inner organs of an organism (Muhr et al., 2023; Purves et al., 2001b). One event critical for organogenesis in vertebrates is neurulation – the process that begins development of the nervous system (Gilbert, 2000b; Purves et al., 2001b).
The vertebrate nervous system is composed of the brain and spinal cord, which make up the central nervous system (CNS), as well as the peripheral nervous system (PNS), which includes all nerves that exist outside of the CNS (Bazira, 2021). In humans, the PNS is split into two subsystems: autonomic and somatic (Bazira, 2021). The autonomic nervous system controls involuntary processes like heartbeat and blood pressure (Bazira, 2021; Gibbons, 2019). Sensory and motor neurons of the somatic PNS, by contrast, are used to carry signals to the brain or carry out commands from the brain, respectively (Bazira, 2021; Ju, 2020). The brain relies on information from the PNS to monitor the internal and external environment and then uses that information to respond to its surroundings. In humans, the enteric nervous system (ENS) is part of the autonomic nervous system in the PNS and innervates the gastrointestinal tract. The ENS acts independently of the CNS, though the ENS and CNS are interconnected through the vagus nerve (Purves et al., 2001a).
However, other animals organize their nervous systems in other ways. Cnidarians have a simple nervous system and are thought to descend from some of the first animals to possess a nervous system, along with Ctenophores (Burkhardt, 2022; Grimmelikhuijzen & Westfall, 1995; Hiroshi Watanabe, Fujisawa, et al., 2009). The basic Cnidarian nervous system organization is that of a nerve net, in which some locations can be condensed to form more complex structures (Grimmelikhuijzen & Westfall, 1995; Hiroshi Watanabe, Fujisawa, et al., 2009). Echinoderms, in contrast, have three main components to their nervous systems—ectoneural, hyponeural, and endoneuroal systems (Cobb, 1987; García-Arrarás et al., 2001). The ectoneural system has both sensory and motor functions and is composed of the nerve ring surrounding the mouth and the radial nerve cords (Cobb, 1987; Formery et al., 2021; García-Arrarás et al., 2001). The hyponeural system is primarily motor and is composed of the radial nerve cords associated with the skeletal muscle system (Cobb, 1987; Formery et al., 2021; García-Arrarás et al., 2001). Less is known about the entoneural system, which is only present in asteroids and crinoids (García-Arrarás et al., 2001; Haugh, 1975; Mercurio et al., 2024).
In vertebrate development, the nervous system begins to form early, starting with a process called neurulation (Colas & Schoenwolf, 2001; Gilbert, 2000b; Purves et al., 2001b). Neurulation in humans begins in weeks 3-4 of gestation (Chhetri & Das, 2023). During neurulation, the ectoderm is induced to become a neural plate following signals from the mesoderm (Chhetri & Das, 2023; Colas & Schoenwolf, 2001; Gilbert, 2000b). The neuroectoderm inverts to give rise to the neural tube and, nearby, the neural crest (Chhetri & Das, 2023; Gilbert, 2000b; Purves et al., 2001b). The neural tube, which provides cells for the central nervous system, is the source of CNS neurons, oligodendrocytes (which myelinate neurons in the CNS), astrocytes, and ependymal cells (Chhetri & Das, 2023; Gilbert, 2000b; Rasband, 2015). In the neurula stage, areas of the neural tube are also further specialized into three distinct regions: the forebrain, midbrain, and hindbrain vesicles (Cowan, 1978, 1979). These three regions give rise to all the structures in the developed brain (Cowan, 1978, 1979). The neural crest gives rise to the PNS as well as a variety of other cell types and tissues, including pigment cells, cartilage, and smooth muscle (Mayor & Theveneau, 2013). The neural crest provides cells for the PNS, including PNS neurons, Schwann cells (which myelinate neurons in the PNS), and other glia (Mayor & Theveneau, 2013; Rasband, 2015). Sensory, motor, and enteric nervous systems each develop from the neural crest through distinct but interrelated processes (Mayor & Theveneau, 2013).
Figure 1
Schematic Overview of Neurulation
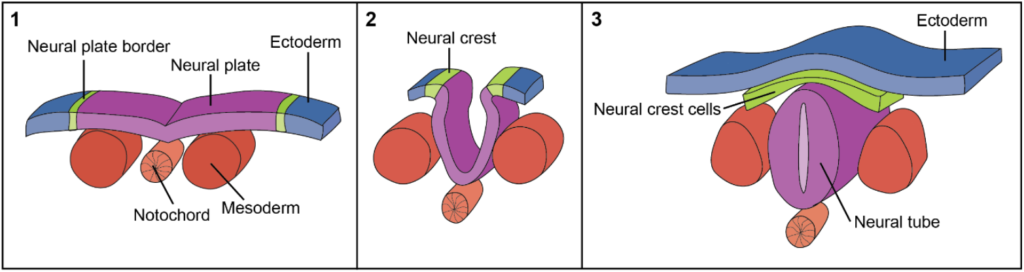
Note. Adapted from Development of the Central Nervous System – Spinal Cord, TeachMeAnatomy.
The nervous system is formed by neural progenitors, also called neuroblasts, that are characterized by their structure, pattern of differentiation, and transcriptional profiles (David J. Anderson, 2001; Le Douarin & Teillet, 1974). Several factors are known to influence neuroblasts to regulate the mature neuron population. After the establishment of the neurectoderm, basic helix-loop-helix transcription factors are upregulated in pro-neural clusters of neurectodermal cells, promoting the developmental program of neuroblasts (Hartenstein & Wodarz, 2013; Huang et al., 2014; Powell & Jarman, 2008; Quan & Hassan, 2005). Conserved mammalian SoxB family transcription factors are critical for defining cell populations that will become neuroblasts (Sasai, 2001). Transcriptional regulators like Six3/6, Pax6, Pax3/7, and Nkx2.1/2.2 are also expressed early and are restricted to specific locations in the neurectoderm. These genes play active roles in triggering neurogenic potential while inhibiting neural differentiation, thereby maintaining each cell’s proliferative state (Arendt et al., 2016; Bylund et al., 2003; Elkouris et al., 2011; Sasai, 2001). Induction and patterning of early neural determinants is achieved through signaling through conserved pathways like Wnt and BMP, as well as their inhibitors (Darras et al., 2011; Marlow et al., 2013; Niehrs, 2010; Sinigaglia et al., 2013). Notch signaling also helps mediate an inhibitory feedback loop that is activated by pro-neural genes, temporally and spatially restricting formation of neural progenitors (Hartenstein & Wodarz, 2013; Kageyama et al., 2009). Neural progenitors proliferate and differentiate variably depending on the type of neural cells they will become and the region of the nervous system in which they will be located.
Further, temporal patterning is required for proper nervous system development (Blackshaw & Cayouette, 2025; El-Danaf et al., 2023). The underlying factors controlling temporal patterning have been best studied in Drosophila (El-Danaf et al., 2023; Grosskortenhaus et al., 2005; Isshiki et al., 2001; Ulvklo et al., 2012). In flies and vertebrates, neuroblasts divide several times asymmetrically to self-renew, to generate intermediate progenitors, and to generate neurons of distinct fates (Doe, 2017; Holguera & Desplan, 2018). Temporal identity factors were first characterized in the Drosophila ventral nerve cord, where sequential expression of hunchback, krüppel, pdm, castor, and grainyhead maps out the temporal steps of neuroblast division and differentiation into specific neuronal subtypes (Grosskortenhaus et al., 2005; Ulvklo et al., 2012). While Drosophila temporal identity factors are conserved in vertebrates, specific roles in regulating the timing of neural development are mostly conserved in the eye (Elliott et al., 2008; Javed et al., 2020, 2023).
After neuronal birth, axons and dendrites must extend out along specific routes to create a network of connections between different parts of the nervous system, and connections must be adjusted and refined (Alberts et al., 2002). Typically, a neuron will have one long axon and several shorter dendrites that project toward target cells, though cell morphology can vary by species and neuronal cell type (Alberts et al., 2002). Extension of axons is driven by a growth cone that is present at the tip of the process (Alberts et al., 2002; Maloney & Bamburg, 2011; Pérez-Ferrer & Herrera, 2025; Sousa & Sousa, 2021). Axons and dendrites connect neurons to other neuronal or non-neuronal cells to form synapses (Alberts et al., 2002; Qi et al., 2022). In development, some organisms create an abundance of synapses during synaptogenesis, then use synaptic pruning to create the highly precise network of connections that is seen in the adult nervous system (Alberts et al., 2002; Faust et al., 2021; Qi et al., 2022; Sakai, 2020; Wolterhoff & Hiesinger, 2024).
Neural Diversity
Through neural development, a broad range of neurons are specified and mature, each with its own structure, function, and connections. Recent studies have shown that the human brain has over 3000 cell types, with each cell type possessing a unique transcriptional identity (Ament et al., 2023; Siletti et al., 2023). Heterogeneity in the brain is in part due to different neuron types, often identified by their neurotransmitters (Cizeron et al., 2020; Que et al., 2019; Sugino et al., 2019). In addition to several different types of neurons, the brain has a highly heterogeneous population of glial cells that vary in location, gene expression, and structure (Bisht et al., 2016; Butt & Verkhratsky, 2018; Khakh & Deneen, 2019; Oberheim et al., 2012; Silvin & Ginhoux, 2018; Stratoulias et al., 2019; Tan et al., 2020; Westergard & Rothstein, 2020). Neural diversity extends beyond the central nervous system into all parts of the peripheral nervous system, with various neuronal and glial cell types found in the nervous system throughout the body.
Recent breakthroughs in bar-coding technologies and single-cell sequencing have allowed for lineage tagging and analysis of gene expression profiles to better understand how cellular diversity arises within the nervous system (Alemany et al., 2018; Bowling et al., 2020; McKenna et al., 2016; Raj et al., 2018). During nervous system development, neural progenitor cells initially divide symmetrically to expand the progenitor pool, followed by frequent asymmetric division to create an intermediate progenitor pool (Ge et al., 2022; Taverna et al., 2014). It is widely accepted that neuronal fate decisions occur at the progenitor level, with more recent evidence suggesting environmental and wiring factors contribute to neuronal identity and diversification post-mitotically (Lodato et al., 2011; Lodato & Arlotta, 2015; Pla et al., 2006; Poliak et al., 2016; Pouchelon et al., 2014). Further, neurogenesis is driven by temporal and spatial patterning, with the possibility that these codes are coordinated to precisely control neuronal production (Briscoe et al., 2000; Sagner & Briscoe, 2019; Telley et al., 2019).
Dopaminergic Neurogenesis
Dopaminergic neurons are specialized neurons located in the CNS and PNS that produce and release the neurotransmitter dopamine (Chinta & Andersen, 2005; Klein et al., 2018). The identification and location of dopaminergic neurons was first confirmed in rodent tissue samples utilizing histofluorescence methods (Falck et al., 1962). Mature dopaminergic neurons are also characterized by their expression of tyrosine hydroxylase (TH), an enzyme involved in the dopamine biosynthesis pathway, though norepinephrinergic neurons also express TH (Blanchard et al., 1993; Moore & Bloom, 1979; Raisman‐Vozari et al., 1991; Walkers & Holden-Dye, 1989). In the human CNS, the main source of dopamine is midbrain dopaminergic neurons, though dopaminergic neurons are also found in the diencephalon and olfactory bulb (Björklund & Lindvall, 1984). In the midbrain, several factors promote the development of dopaminergic neurons from stem cells and early progenitors. Sonic hedgehog (Shh) and fibroblast growth factor 8 (FGF8) signals provide essential cues for early dopaminergic progenitor cells (Hynes & Rosenthal, 1999; Rosenthal, 1997). Several early transcription factors help regulate early patterning in the ventral midbrain, including basic Helix-Loop-Helix transcription factors Hes1 and Nato3 (Kameda et al., 2011; Ono et al., 2010). Homeodomain transcription factors like Lmx1a, Lmx1b, and Msx1 help establish the identity of ventral midbrain dopamine neural precursors (Andersson et al., 2006; Failli et al., 2002; Smidt et al., 2000). Ngn2 and Nurr1 are then expressed in more specialized cells that go on to become dopaminergic neurons (Andersson et al., 2006; Hyun Jung Kim et al., 2007; Law et al., 1992; Zetterström et al., 1996).
All dopaminergic neurons in the PNS, including those in the ENS, arise from the neural crest (R. B. Anderson et al., 2013; Hao et al., 2013; Mayor & Theveneau, 2013). While little work has been done to explore the development of dopamine-producing cells in the non-enteric PNS, we do know that dopamine in the body periphery originates from a few different sources: noradrenergic neuronal fibers; neuroendocrine cells; and the adrenal medulla (Fan & Katz, 1993; Goldstein & Holmes, 2008; Katz et al., 1983; Pearse, 1969; Wahbe et al., 1982). ENS neurons come from precursors derived from the neural crest known as enteric neural crest-derived cells (ENCDC) (Le Douarin & Teillet, 1973, 1974; Yntema & Hammond, 1954, 1955). Several factors are required for ENCDCs to enter the gut and migrate through the entire organ, including PHOX2B, RET, and SOX10 (Jaesang Kim et al., 2003; Z. S. Li et al., 2004; Marcos & Pachnis, 1996; Pattyn et al., 1999; Taraviras et al., 1999). SOX10 also works with ZEB2 to maintain the potency of ENCDCs and inhibit neuronal differentiation through EdnrB signaling (Jaesang Kim et al., 2003; Yuli Watanabe et al., 2017). Ascl1 is critical for both enteric gliogenesis and enteric neurogenesis, specifically for dopaminergic neurons (Memic et al., 2016).
In addition to vertebrate dopaminergic neurogenesis, research in invertebrates has provided insights into the mechanisms through which dopaminergic neurons arise during development. For example, identification and lineage tracing of heterogeneous populations of dopaminergic neurons in both embryonic and larval stages of Drosophila melanogaster have been completed (Hartenstein et al., 2016). While many factors required for dopaminergic neuron development are conserved, Drosophila dopaminergic neurons also require expression of genes like sim, wingless, and gooseberry, which encode transcription factors (sim and gooseberry) or Wnt family members (wingless) (Bhat, 2007; Duman-Scheel et al., 1997; X. Li & Noll, 1993; Nambu et al., 1990; Richter et al., 1998; Skeath et al., 1995).
Further, hermaphroditic Caenorhabditis elegans have 302 neurons, eight of which are dopaminergic (J. Sulston et al., 1975; White et al., 1986). Of these eight dopaminergic neurons, there are four different classes that arise at different times during development, are located in different regions of the nervous system, and come from distinct cell lineages (J. E. Sulston et al., 1983; J. E. Sulston & Horvitz, 1977; White et al., 1986). Though these populations of neurons are distinct, several factors impact all 8 C. elegans dopaminergic neurons, including AST-1 (an ETS domain transcription factor), CEH-43 (a distal-less ortholog), and several Pbx genes (Doitsidou et al., 2008; Flames & Hobert, 2009; Siehr et al., 2011). Lin-32, which is a basic helix-loop-helix transcription factor, and a PAX6 homolog also regulate dopaminergic neuron identity and cell number, respectively (Doitsidou et al., 2008, 2013). Because C. elegans has distinct dopaminergic neuron populations, there are likely factors that are necessary for specific sets of dopaminergic neuron populations. At least one such factor has been identified. ztf-6 is a gene encoding a zinc finger transcription factor that regulates dopaminergic neurons in a lineage-specific manner, impacting distinct dopamine neuron-producing lineages differently (Doitsidou et al., 2018).
Dopaminergic Neurons & Parkinson’s Disease
Loss or dysfunction of dopaminergic neurons is associated with neurological diseases and disorders like Alzheimer’s Disease (Nobili et al., 2017), Huntington’s Disease (Bédard et al., 2011; Rangel-Barajas et al., 2015), Multiple Sclerosis (Rangel-Barajas et al., 2015), and most commonly Parkinson’s Disease (Bédard et al., 2011; Damier et al., 1999; Hirsch et al., 1988; Parkinson, 2002). Parkinson’s Disease is a neurodegenerative disease that impacts motor and non-motor systems through loss of dopaminergic neurons in a section of the midbrain, the substantia nigra, that provides dopamine to the basal ganglia (Damier et al., 1999; Hirsch et al., 1988). Motor dysfunction is one of the earliest symptoms, with patients exhibiting tremors, hypokinesia, and rigidity (Parkinson, 2002). Parkinson’s Disease was initially thought to be a sporadic neurodegenerative disorder that had greater prevalence depending on certain environmental factors; genetic mutations have been identified in some familial forms of Parkinson’s Disease (Duvoisin, 1984; Lesage & Brice, 2009). The first gene mutation associated with Parkinson’s Disease was a mutation in the alpha-synuclein (SCNA) gene (Polymeropoulos et al., 1997). One pathological hallmark of Parkinson’s Disease is the presence of alpha-synuclein aggregates in Lewy bodies or Lewy neurites in affected neurons. Aggregates also include other components like phosphorylated neurofilaments and ubiquitin (Srinivasan et al., 2021). The genetic mutation that conveys the greatest risk of Parkinson’s Disease occurs in a gene that encodes a lysosomal enzyme, glucocerebrosidase (GBA) (Sidransky et al., 2009; Sidransky & Lopez, 2012). Other genes implicated in Parkinson’s Disease include LRRK2 and parkin (or PARK2), which contribute to the most common forms of inherited Parkinson’s Disease (Matsumine et al., 1998; Paisán-Ruíz et al., 2004; Zimprich et al., 2004). While there have been several genetic risk factors identified for Parkinson’s Disease, the vast majority of cases (85%) arise sporadically with no known hereditary cause (Matsumine et al., 1998; Paisán-Ruíz et al., 2004; Polymeropoulos et al., 1997; Sidransky et al., 2009; Sidransky & Lopez, 2012; Srinivasan et al., 2021; Trevisan et al., 2024; Zimprich et al., 2004). In these cases, environmental toxins and brain trauma may contribute to some, but not all, sporadic cases.
Many neurodegenerative diseases, like Parkinson’s Disease and Amyotrophic lateral sclerosis, have been associated with the accumulation of protein aggregates (e.g., alpha-synuclein aggregates in Parkinson’s Disease) in degenerating and dying neurons (Koszła & Sołek, 2024; Polymeropoulos et al., 1997; Sweeney et al., 2017). Whether these protein aggregates are causative or correlated with neuronal decline remains controversial, but the evidence showing the presence of abnormal protein accumulation across neurodegenerative diseases is indisputable. Cells mount defenses to help fix proteostasis problems that may arise, including protein dysfunction and misfolding (Barmaki et al., 2023; Díaz-Villanueva et al., 2015). Neuronal chaperones are proteins that aid in several processes to help prevent neuronal death due to protein aggregation (Ellis, 1988; Hartl et al., 1992; Hartl, 1996). Chaperones assist in the folding of nascent or misfolded proteins into their correct three-dimensional structures and bind to misfolded proteins to prevent harmful aggregation. When chaperones are not successful, misfolded proteins are targeted to the ubiquitin-proteasome system for degradation (Satapathy & Wilson, 2022; Heather L. Smith et al., 2015). Because of their neuroprotective roles, neuronal chaperones and components of the ubiquitin-proteasome system could be a target for treatment of neurodegenerative diseases (Luo et al., 2010; Sharma et al., 2024). Increasing activity of key proteostasis factors could slow disease progression.
Parkinson’s Disease is most commonly attributed to defects in the central nervous system, but dopaminergic neurons also exist outside the CNS in the peripheral nervous system, including the enteric nervous system (Chalazonitis et al., 2022; Chinta & Andersen, 2005). The majority of research on Parkinson’s Disease has focused on loss of dopamine in the CNS, specifically the substantia nigra (Barker et al., 2017; Björklund & Stenevi, 1979; Brundin et al., 1988; Damier et al., 1999; Freed et al., 2001; Nagarajan et al., 2014; Yang et al., 2008). However, as early as 1931, researchers noticed Lewy bodies in neurons outside of the CNS (den Hartog Jager & Bethlem, 1960; Hechst & Nussbaum, 1931). Braak’s 6-stage theory proposes that Parkinson’s Disease-associated neurodegeneration begins in the PNS (specifically the ENS) and progresses to the CNS through the vagus nerve (Braak et al., 2003; Braak & Del Tredici, 2017). The Braak staging system is based on the progression of alpha-synuclein structures in over 100 Parkinson’s Disease patients presenting with alpha-synuclein aggregates (Braak et al., 2005, 2003). According to this model, the disease begins in the dorsal motor nucleus of the vagus nerve, eventually progressing and extending into and throughout parts of the brain (Braak et al., 2005, 2003). Though the clinical progression of Parkinson’s Disease cannot yet be fully explained, data support a spread of the disease between different parts of the nervous system. For example, several studies demonstrated accumulation of alpha-synuclein in the enteric nervous system before the CNS (Iranzo et al., 2014; Kupsky et al., 1987; Qualman et al., 1984; Wakabayashi et al., 1988).
Parkinson’s Disease has been studied for over a century, yet there is no cure, and treatment options for Parkinson’s Disease revolve around managing symptoms through drugs that help increase dopamine synthesis and uptake. Levodopa has been considered the “gold standard” for treating symptoms of Parkinson’s Disease, however long-term use of the drug, which is the immediate precursor for dopamine synthesis in the substantia nigra, causes adverse side effects (Fahn, 2005; Grandas et al., 1998; Melamed, 1979; Salat & Tolosa, 2013; Sethi, 2010; Tolosa et al., 1975). Alternatively, dopamine agonists can be used to activate dopamine receptors, but they also have negative side effects (Calne et al., 1974; Cotzias et al., 1970). In the past several decades, research on stem cell-based therapies emerged, with stem cells posing an attractive treatment option for introducing functioning dopaminergic neurons in affected areas. Dopaminergic neuron transplants in animal models began as early as the 1970s, with transplantation of rat fetal neurons into injured rodents (Björklund et al., 1976; Björklund & Stenevi, 1979). Pioneering work demonstrated that fetal dopamine neurons from rodents could be incorporated into a dopamine-deficient rodent brain and restore dopamine pathways, paving the way for experiments transplanting human fetal neurons in the same manner (Björklund et al., 1980, 1981; Ungerstedt et al., 1974). Grafting human fetal brain tissue into rodent models allowed functional recovery, though development and innervation of these cells took longer than the previously tested rodent cells (24 weeks opposed to 8 weeks) (Brundin et al., 1988). Not only are human dopamine neurons able to functionally recover in the immediate location of the graft, but they also extend axons over long distances to relevant cortical areas and the striatum of the transplanted rodent (Grealish et al., 2014; Moriarty et al., 2022; Wictorin et al., 1992; Xiong et al., 2021).
Early clinical trials based on grafting human fetal ventral mesencephalic tissue into human patients reported variable results and no significant changes in Parkinson’s Disease patients, especially for recipients over 60 years old (Freed et al., 2001; Olanow et al., 2003). Further, a few subjects who received grafts were examined 11-24 years later, and researchers found that grafted dopamine neurons displayed alpha-synuclein pathology similar to diseased neurons (Kordower et al., 2008; Jia Yi Li et al., 2008). The presence of Lewy bodies in transplanted cells suggested that Parkinson’s Disease can propagate from host cells to grafted cells, though the majority of the grafted cells were unimpaired after over a decade (Kordower et al., 2008; Jia Yi Li et al., 2008).
In the 2010s, there were breakthroughs in human pluripotent stem cell therapies, including human embryonic stem cells (Thomson, 1998) and human induced pluripotent stem cells (Takahashi et al., 2007). The first protocols for generating dopamine neurons from human pluripotent stem cells were adapted from mouse protocols and included induction of neural differentiation and exposure of cells to morphogens and growth factors like Shh, FGF8, brain-derived neurotrophic factor, and glial cell line-derived neurotrophic factor (Sánchez-Pernaute et al., 2001; Sonntag et al., 2007; Zhang et al., 2001). Surprisingly, embryonic stem cell- and induced pluripotent stem cell-derived dopaminergic neurons performed poorly post transplantation and midbrain identity was not confirmed (Jong Hoon Kim et al., 2002; Sánchez-Pernaute et al., 2001; Yang et al., 2008). Because midbrain dopamine neurons arise from floorplate progenitors marked by coexpression of FOXA2 and LMX1A, alternate protocols were then developed (Bonilla et al., 2008; Joksimovic et al., 2009; Ono et al., 2007). These adjustments, in combination with SMAD inhibition, early exposure to Shh, and Wnt pathway activation, allowed for the development of midbrain dopamine neurons that could functionally replace disease-impacted cells after transplantation (Grealish et al., 2014; Kirkeby et al., 2012; Kriks et al., 2011). Clinical trials for dopamine cell therapy in humans began in the late 2010s and early 2020s, with two groups recently publishing results from their Phase I and/or Phase II trials (Sawamoto et al., 2025; Tabar et al., 2025). Emerging data suggest that dopaminergic cell therapies derived from either human pluripotent stem cells (Sawamoto et al., 2025) or human embryonic stem cells (Tabar et al., 2025) could be safe and could produce clinical benefits for Parkinson’s Disease, including improved motor symptoms (Sawamoto et al., 2025; Tabar et al., 2025). While treatments for Parkinson’s Disease have improved over the past five decades, the most recent trials still indicate modest benefits, and long-term studies will still need to confirm the efficacy and persistence of cell-based therapies. Thus, additional interventions will likely be needed for a Parkinson’s Disease cure.
Regenerative Capabilities Across the Animal Kingdom
One possible approach to functionally restore cells in the case of nervous system injury could be inducing new production of neurons in the brain to replace diseased and dying cells. The ability to regenerate cells is common throughout non-human members of the animal kingdom, with many species repairing and healing neural tissue after injury through limited cell birth or de/transdifferentiation (Arenas Gómez et al., 2020; Iismaa et al., 2018; Tanaka & Reddien, 2011). Most mammals regenerate poorly, but some mammals are capable of regeneration limited to some tissues or developmental stages (Iismaa et al., 2018; Storer & Miller, 2020). For example, humans regenerate parts of organs like the liver, skin, and uterine lining (Cousins et al., 2022; Ferenczy et al., 1979; Gargett & Ye, 2012; He et al., 2021; Rowlatt, 1979; Takeo et al., 2015; Wei et al., 2021). However, there are limitations to human regenerative ability, especially when it comes to the nervous system (Iismaa et al., 2018; Varadarajan et al., 2022). While humans can repair the peripheral nervous system to an extent, the ability to recover after central nervous system injury is extremely limited (Contreras et al., 2022; Gordon, 2020; Varadarajan et al., 2022; Widodo et al., 2025). Thus, brain injuries and diseases of the CNS are often incurable. Treatments for these debilitating diseases focus on prolonging life, rehabilitation, and palliative care or symptom reduction rather than a true cure (Fahn, 2005; Grandas et al., 1998; Shusharina et al., 2023; Tolosa et al., 1975).
In contrast, some remarkable animals complete successful regeneration throughout the nervous system. Axolotls and zebrafish regenerate cells within the brain and spinal cord, though neither organism can fully reestablish perfect neuronal diversity and connectivity after injury (Amamoto et al., 2016; Caldwell et al., 2019). Zebrafish utilize a population of resident progenitors that give rise to mature cells in the locations where they are needed after injury (Grandel et al., 2006). While zebrafish and axolotls do possess incredible regenerative capabilities, the ability to regrow an entire nervous system de novo after injury is rare and limited to a small number of animals, including Hydra (a cnidarian) and planarian flatworms (Umesono & Agata, 2009; Hiroshi Watanabe, Hoang, et al., 2009). To answer questions specific to successful, whole-brain regeneration, I sought to use planarians due to their robust regeneration of the entire body, including all parts of the nervous system.
Planarians As a Model for Regeneration
Planarians are non-parasitic, freshwater flatworms of the phylum Platyhelminthes that are known for their remarkable regenerative capabilities. Though several species of planarians exist, scientists most often use two species to study regeneration: Dugesia japonica and Schmidtea mediterranea. I used the asexual strain of S. mediterranea to answer questions about how successful nervous system regeneration occurs. S. mediterranea undergoes robust, whole-body regeneration and replenishes all tissues after nearly any injury, including the creation of the nervous system de novo (Inoue et al., 2004; Reddien & Sánchez Alvarado, 2004; Ross et al., 2017). Planarians accomplish great regenerative feats through a population of adult pluripotent stem cells, called neoblasts, that give rise to all mature cell types in the planarian body (Baguñà et al., 1989; Morita et al., 1969; Wagner et al., 2011a; Wenemoser & Reddien, 2010). Stem cells produce new cells that differentiate to populate all mature organs in the body, as well as the connective tissue. Planarian pluripotent stem cells are found throughout the body and are only absent from the most anterior part of the head and the pharynx, the feeding organ, which are the only two parts of the animal that cannot regrow the rest of the body when isolated (Reddien et al., 2005).
Pluripotent stem cells make up approximately 20% of all cells within the planarian body and respond to signals after injury (Hayashi et al., 2006; Reddien et al., 2005; Wagner et al., 2011b). In the instance of lost tissue, stem cells respond by dividing and moving to the location of injury to replenish lost cells and tissue (Baguñà et al., 1989; Takeda et al., 2009; Wagner et al., 2011b; Wenemoser & Reddien, 2010). Planarians do not have resident progenitor cells within organs, so it is necessary for stem cells and their progeny to migrate to locations where they are needed. Planarian pluripotent stem cells respond to injury with two proliferative bursts, the first of which occurs at 6 hours and the second at 48 hours post-injury (Baguñà, 1976; Saló & Baguñà, 1984; Wenemoser & Reddien, 2010). Pluripotent stem cells divide to replenish the stem cell population and to create cells that differentiate and eventually mature into specified cell types.
Planarian pluripotent stem cells, or neoblasts, were originally thought to be a relatively homogeneous population of cells defined by morphological characteristics like a large nucleus-to-cytoplasmic ratio, expression of Smedwi-1, and location in the mesenchymal tissue (Baguñà, 2012; Baguñà et al., 1989; Reddien et al., 2005). However, many studies have shown that the neoblast population is heterogeneous and that many neoblasts express specific markers of restricted cell fates (Molinaro & Pearson, 2016; Raz et al., 2021; van Wolfswinkel et al., 2014). Pluripotent neoblasts capable of repopulating the animal’s tissue and giving rise to all cell types are termed clonogenic neoblasts, though this is a functional definition rather than a molecular one (Reddien et al., 2005). Four subpopulations have been identified that express Smedwi-1 as well as transcription factor-encoding genes that are tissue-specific. These neoblast populations—ζ-, σ-, γ-, and ν-neoblasts—are thought to be specialized progenitors with narrower fate (van Wolfswinkel et al., 2014). ζ-neoblasts are considered specialized cells that help maintain and give rise to the epidermal lineage, and they express high levels of genes like egr-1, fgfr-1, soxP-3, and zfp-1 (van Wolfswinkel et al., 2014). σ-neoblasts have a broad role in producing cells of several different lineages, including other neoblast types, like ζ-neoblasts (van Wolfswinkel et al., 2014). σ-neoblasts are characterized by high expression of genes such as soxP-1, soxB-1, fgfr-4, and nlk-1 (van Wolfswinkel et al., 2014). In this same study, a third population of neoblasts, considered to be a subclass of σ-neoblasts, was identified: γ-neoblasts. This population is likely an intestine-specific progenitor pool, and it is characterized by elevated expression of genes like gata4/5/6, hnf4, and nkx2.2 (van Wolfswinkel et al., 2014). Finally, a fourth population of neural-specific progenitors was identified and termed the ν-neoblasts. This neural-committed population of neoblasts expresses high levels of genes, including ston-2, elav-1, and ptprd-9 (Molinaro & Pearson, 2016). Importantly, recent work indicates that stem cells with lineage-specific markers retain their potency and can return to a less specialized state after division, suggesting that pluripotency is not linked to a unique stem cell class in planarians (Raz et al., 2021). Additionally, work continues to reveal factors necessary for stem cell maintenance and function, as well as signals that tell a stem cell to differentiate and specialize into each mature cell type (Benham-Pyle et al., 2023; King et al., 2024).
Planarian Organ Systems
Planarians have complex organ systems, including nervous, muscle, digestive, epidermis, excretory, and reproductive systems that are held together by connective tissue called the parenchyma (Roberts-Galbraith & Newmark, 2015) (Figure 2A-D). For this work, I focus on the planarian nervous system, which is made up of a central nervous system, a peripheral nervous system found throughout the body, and a pharyngeal nervous system (PhNS) found in the planarian feeding organ (Roberts-Galbraith & Newmark, 2015; Ross et al., 2017) (Figure 2A-B). The central nervous system is composed of a bi-lobed brain, two nerve cords on the ventral side of the animal, and two eyespots on the dorsal side of the planarian (Agata et al., 1998; Carpenter et al., 1974; Krugelis MacRae, 1964) (Figure 2A). The brain also has brain branches that reach out laterally from the brain and relay chemosensory signals (MacRae, 1967) (Figure 2A). The peripheral nervous system is composed of nerve plexuses that sit between and around different organ systems (Figure 2B). The subepidermal nerve plexus innervates the space between the epidermis and the body wall musculature, and likely functions in sensory processes or in the regulation of epidermal motile cilia (Baguñà & Ballester, 1978) (Figure 2B). The submuscular nerve plexus is located below the muscle wall, and the gastrodermal nerve plexus surrounds the intestine (Baguñà & Ballester, 1978) (Figure 2B). The pharyngeal nervous system, composed of two ring-like nerve plexuses, is found solely within the pharynx and is important for pharyngeal movement in feeding behavior. While characterization of the CNS in planarians has been a focus of the field, the PNS remains poorly characterized, with sparse molecular and genetic information about the identities and developmental processes for PNS or PhNS neurons. The planarian nervous system contains dozens of neuron types, which have been defined by their gene expression and—less commonly—function. Planarian neurons produce several different neurotransmitters including acetylcholine, dopamine, GABA, octopamine, serotonin, norepinephrine, and over 50 neuropeptides (Clay et al., 2025; Collins et al., 2010; Nishimura et al., 2010; Nishimura, Kitamura, Umesono, et al., 2008; Nishimura, Kitamura, Inoue, et al., 2008; Nishimura, Kitamura, Inoue, Umesono, Sano, et al., 2007; Nishimura, Kitamura, Inoue, Umesono, Yoshimoto, et al., 2007; Ong et al., 2016).
Figure 2
Planarian Organ Systems
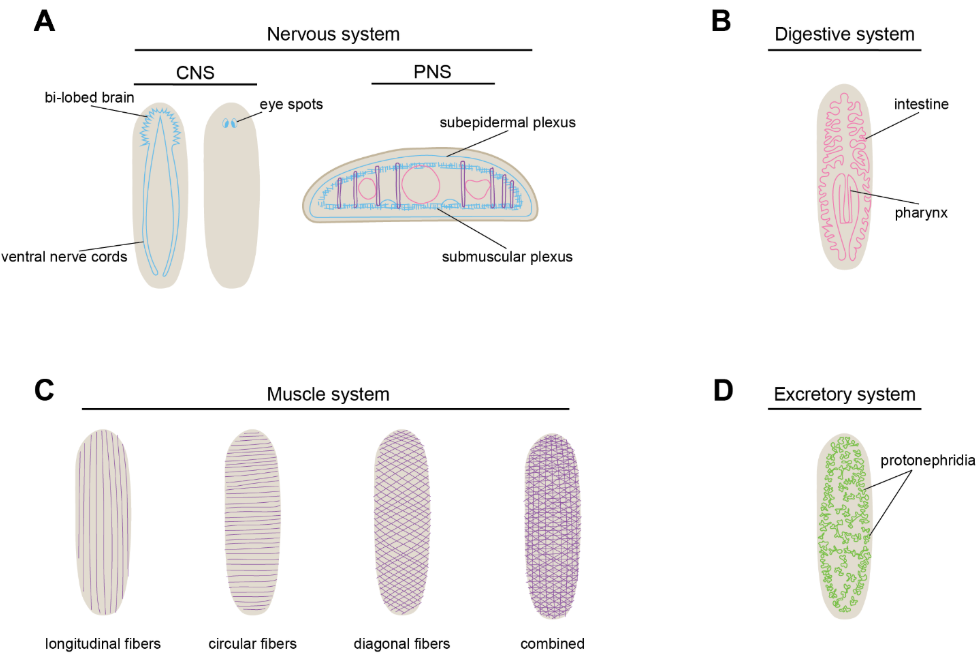
Notes. A) Diagram of the planarian nervous system. Central nervous system (CNS) on left, peripheral nervous system (PNS) on right. B) Diagram of the digestive system. C) Diagram of planarian musculature. D) Diagram of the planarian excretory system.
While we do not know the entire mechanism through which any neuron type is specified and differentiated, we do know some key regulators of the planarian nervous system. For example, soxB1-2, coe, and sim are required for broad nervous system regeneration (Cowles et al., 2013, 2014; Ross et al., 2017). ets-1 and hedgehog signaling are required for proper regeneration and patterning of planarian glia, neuronal support cells (Chandra et al., 2023; Wang et al., 2016). Finally, there has been substantial work to uncover mechanisms that allow specification and maturation of serotonergic neurons, including identification of pitx and lhx1/5-1 as terminal differentiation factors for neurons producing serotonin (Currie & Pearson, 2013; März et al., 2013). Importantly, additional factors required for determining the exact spatial and regional localization of neurons, and establishing neurons in the correct numbers, types, and ratios, have yet to be elucidated. Uncovering the mechanisms required for proper neuronal regeneration in the correct contexts for a single mature neuron type will likely provide critical information for differentiation pathways for other mature cell types in the planarian.
Within the nervous system, we do know the entire pathway from a pluripotent stem cell to mature eye cells, including pigment cup cells and photoreceptor cells (Atabay et al., 2018; Emili et al., 2019; Lapan & Reddien, 2011, 2012a). Planarians have two eyespots with mature cells, and a trail of eye progenitors that lead toward them (Agata et al., 1998; Carpenter et al., 1974; Inoue et al., 2004; Lapan & Reddien, 2011, 2012b). To determine the order in which transcription factors need to be active for proper eye formation, scientists completed a combination of fluorescent in situ hybridization experiments labeling stem cells, mature eye cells, and transcription factor-encoding mRNAs that were thought to act in the process (Lapan & Reddien, 2011). Transcription factor-encoding genes expressed with the stem cell marker were determined to mark early progenitors, and those expressed with the mature cell type marker were determined to act later in the lineage (Lapan & Reddien, 2011).
We also know the entire pathway from a planarian pluripotent stem cell to mature epidermal cells, and we understand many of the factors at work in between (Eisenhoffer et al., 2008; Tu et al., 2015; Zhu & Pearson, 2016). The combination of 5-bromo-2′-deoxyuridine labeling with in situ hybridization experiments allowed lineage tracing in planarians and created a guide for determining lineage for other cell types as well (Eisenhoffer et al., 2008). Other factors have been identified in pathways of specification for different mature cell types in the planarian, including muscle, gut, and protonephridia. Proper expression of genes such as myoD, foxF-1, nk4, gata4/5/6-2, and gata4/5/6-3 is required for proper planarian muscle regeneration (Cebrià, 2016; Lucila Scimone et al., 2017; Scimone et al., 2018). Regeneration and/or maintenance of cells in the intestine requires proper expression of genes like gli-1, RREB2, lhx2/9-1, lhx1/5-2, LDB1, SSDP2, and gata4/5/6 (Flores et al., 2016; Forsthoefel et al., 2020; Medlock-Lanier et al., 2024; Molina et al., 2023). The planarian excretory system is governed by transcription regulators like six1/2-2, hunchback, eya, sall, and POU2/3 (Scimone et al., 2011). Additionally, an EGF receptor, EGFR-5, is required for proper branching morphogenesis and maturation of planarian protonephridia (Rink et al., 2011).
Planarian Polarity and Body Plans
In addition to replenishing all missing cells after injury, planarians faithfully reproduce the body plan with predictable patterning, using critical polarity signals that govern the body axes. Planarians have anterior-posterior, dorsal-ventral, and medial-lateral axes that are governed by distinct mechanisms. In planarians, the anterior-posterior axis is governed by posterior Wnt signaling proteins, similar to other metazoans (Hardin & King, 2008; Logan & Nusse, 2004; Marikawa, 2006; Schier & Talbot, 2005). β-catenin functions downstream of canonical Wnt signaling and inhibits anterior pole identity. Knockdown of β-catenin through RNA interference (RNAi) results in ectopic head growth and regeneration (Gurley et al., 2008; Iglesias et al., 2008; Petersen & Reddien, 2008), indicating a negative regulation of brain regeneration.
The planarian dorsal-ventral axis is governed by bone morphogenetic protein (BMP) and anti-dorsalizing morphogenetic protein (ADMP) signals. bmp4 is expressed dorsally and represses ventral admp (Gaviño & Reddien, 2011). In planarians, bmp4 is expressed in clusters of cells along the dorsal midline, as well as a small number of neurons in the brain and ventral nerve cords (Molina et al., 2007; Orii et al., 1998). After knockdown of bmp4, planarians experience ectopic eye expression, indented blastemas, and ultimate failure of regulation at the dorsal midline (Molina et al., 2007; Reddien et al., 2007). bmp4(RNAi) also causes increased admp expression, leading to the conclusion that bmp4 regulates admp through inhibitory mechanisms (Gaviño & Reddien, 2011). admp is expressed in cells along the ventral midline and cells at the lateral edges of the animal (Gaviño & Reddien, 2011). admp(RNAi) animals form indented blastemas and decreased regenerative abilities after a sagittal amputation. Knockdown of admp also led to decreased bmp-4 expression, suggesting that admp activates expression of bmp-4 (Gaviño & Reddien, 2011). The dorsoventral regulators bmp and admp are also regulated by several other factors, including an expanded noggin family (Molina et al., 2007; Ogawa et al., 2002; William C. Smith & Harland, 1992).
Planarians’ bodies are also organized along a medial-lateral axis. The medial-lateral axis is regulated by slit at the midline and lateral Wnt5 through reciprocal patterning (Cebrià et al., 2007; Gurley et al., 2010). slit is expressed along the midline of both the dorsal and ventral sides of the planarian (Cebrià et al., 2007), and knockdown of slit causes misexpression of wnt5 into the planarian midline (Gurley et al., 2010). In contrast, wnt5 is expressed lateral to the ventral nerve cords and peripherally around the planarian body (Gurley et al., 2010). wnt5(RNAi) leads to expression of slit outside the midline and towards the lateral edges of the animal (Gurley et al., 2010).
Planarians faithfully regenerate their entire body, including their complex nervous system, after nearly any injury. Not only do planarians make the right cell types, but they do so in the correct locations, patterning, and ratios. Because of their remarkable regenerative capabilities, planarians are an ideal model with which to uncover mechanisms underlying successful nervous system regeneration. Revealing the factors required for regeneration and maintenance of their complex nervous system will not only further elucidate how planarians regrow without consequence, but could also inform research into therapies for neurological conditions, including brain injury and neurodegenerative disease.
Specification of Dopaminergic Neurons in Planarians
One of the larger goals of my thesis work was to determine the entire genetic pathway from a pluripotent stem cell to a mature dopaminergic neuron in planarians. I proposed a model in which combinatorial mechanisms specify regional location and neurotransmitter identity of dopaminergic neurons concurrently (Clay et al., 2025). However, there is likely some sort of hierarchy to dopaminergic neuron specification, even if there is overlap in the transcriptional regulation. In an effort to parse out the order in which transcription factors are turned on to create dopaminergic neurons in the peripheral nervous system, orthogonal approaches could be used to determine whether fli1-2, irx-4/6, and soxB1-2 are expressed in the early or late progenitors. I found that fli1-2, irx-4/6, and soxB1-2 are all co-expressed with a broad stem cell marker, smedwi-1, and are therefore all expressed in at least some progenitor cells (Clay et al., 2025; Eisenhoffer et al., 2008). In irradiated animals, less specialized stem cells will go away first, followed by early progenitors, late progenitors, and finally mature cell types (Eisenhoffer et al., 2008). Looking at the expression of fli1-2, irx-4/6, and soxB1-2 in irradiated animals over a seven-day timeline will show which of these populations dies off first, providing insight into which of these transcription factor-encoding genes is expressed earliest within progenitors.
In addition to determining the order of expression of the genes I have identified, there are likely other genes that are critical for regeneration and maintenance of dopaminergic neurons in planarians. My current hypothesis is that the genes I have identified are likely enriched for the middle of the specification pathway for dopaminergic neurons. Utilizing available single-cell transcriptomic data, it will be interesting to examine transcripts that are expressed in predicted early neural progenitor populations, especially those that express high levels of smedwi-1 mRNA (Fincher et al., 2018). Another reverse genetic screen examining transcription factors at this stage could provide insight into the earliest fate choices in the dopaminergic neuron pathway. Further, the bulk RNA sequencing I completed will provide information as to what genes may be working downstream in the pathway of dopaminergic neuron specification. Conducting a smaller-scale screen using the bulk RNA sequencing data will help identify additional genes required for the maintenance and maturation of dopaminergic neurons.
Factors Influencing Planarian Neuronal Placement
In my thesis work, I proposed a model in which combinations of transcription factor-encoding genes help specify dopaminergic neurons, both in their neurotransmitter identity and in their regional location (Clay et al., 2025). However, there are likely other factors at play that help regulate both processes. One example is the consistent and robust body-patterning system that planarians possess (Reddien, 2018). Planarians have anterior-posterior, dorsal-ventral, and medial-lateral axes that are strictly regulated (Reddien, 2018). IRX family genes play conserved roles in vertebrates and invertebrates in mediating the dorsoventral axis (Glavic et al., 2002; Gómez-Skarmeta & Modolell, 2002; Itoh et al., 2002; Lecaudey et al., 2004). Interestingly, planarian irx-like has a polarized expression pattern and is only expressed on the ventral side of the animal. This raises the question of whether there are distinct factors regulating dopaminergic neurons in the dorsal and ventral PNS. IRX-like is a likely candidate in helping regulate ventral dopaminergic neurons, but how is this influenced by polarity signaling? Several factors have been identified as regulating polarity axes in planarians (Cebrià et al., 2007; Gaviño & Reddien, 2011; Gurley et al., 2008, 2010; Iglesias et al., 2011; Molina et al., 2007; Ogawa et al., 2002; Orii & Watanabe, 2007; Petersen & Reddien, 2008). Further characterizing the polarity of PNS cells will likely determine other factors impacting dopaminergic neuron fate and regionalization.
Planarians As a Future Model for Neurodegeneration
Unfortunately, there are currently no cures for neurodegenerative diseases, and treatment options available are used to manage symptoms (Cascione et al., 2020; Durães et al., 2018). Several models are being used to model neurodegeneration and identify potential pathways for investigation, including in vivo, in vitro, and in silico models (Dawson et al., 2018; Jones et al., 2022; Valadez-Barba et al., 2020). While these models have provided critical information in understanding the molecular mechanisms behind neurodegenerative diseases, with some studies resulting in clinical trials, more progress is needed. I reasoned that the use of another model system that already possesses perfect regenerative capabilities, including that of its complex nervous system, might provide complementary information and could reveal new molecular mechanisms underlying neurodegeneration.
One approach I employed to model neurodegeneration in planarians was by identifying and characterizing planarian TAR DNA-binding proteins (TARDBPs). Protein aggregates are implicated in several neurodegenerative diseases, including Parkinson’s Disease, Alzheimer’s Disease, and amyotrophic lateral sclerosis (ALS) (Koszła & Sołek, 2024; Polymeropoulos et al., 1997; Sweeney et al., 2017). TDP-43 aggregates are specifically implicated in sporadic and genetic forms of ALS (Sreedharan et al., 2008; Suk & Rousseaux, 2020). However, the exact cause of neuron death after TDP-43 aggregation has yet to be elucidated. In one model, neurons with TDP-43 protein aggregates die due to toxicity when the protein becomes mislocalized and aggregated. In an alternate model, cellular health is primarily compromised due to the loss of healthy TDP-43 protein function. Planarians present a unique organism with which to study the cellular roles of TARDBP because planarians have five homologs of TDP-43. In other organisms with only one TDP-43 homolog, like mice, loss of TDP-43 is embryonic lethal (Kraemer et al., 2010). Because planarians have five TDP-43 homologs, we are able to knock down individual isoforms in the adult animal to examine the subfunctionality of TDP-43. Indeed, I was able to knock down each planarian homolog of TDP-43 and assess whether regeneration of the brain was impaired. I found that tdp-4 is required for proper planarian brain regeneration. Loss of planarian tdp-4 eventually led to animal death through lysis, which led me to examine whether tdp-4 regulated planarian stem cells. I found that tdp-4 causes a reduction in planarian stem cell abundance compared to controls, and I concluded that planarian TARDBPs regulate both brain regeneration and stem cell maintenance.
Currently, most planarian researchers induce regeneration through tissue amputation or other drastic injury. However, this is not physiologically relevant for modeling neurodegeneration. I hypothesized that I might be able to model neurodegeneration in planarians in a manner that is more physiologically relevant to humans. I started by looking at the literature where dysfunction in neuronal chaperones and proteasomal subunits has been implicated in neurodegenerative disease (Ellis, 1988; Hartl et al., 1992; Hartl, 1996; Satapathy & Wilson, 2022) (Figure 3). Using available single-cell transcriptomic data, I identified planarian homologs of chaperones and proteasomal subunits with enriched expression in the nervous system (Fincher et al., 2018). I narrowed this list down to twenty transcripts (e.g., heat shock factor, E3 ubiquitin ligase), which could be used for an RNAi screen to assess their impact on maintenance of brain size and function. Next, I would suggest investigating whether these RNAi conditions lead to neuronal death via protein aggregation. Utilizing TUNEL to examine cell death by apoptosis could help address the first possibility, and I have worked to develop this protocol in the lab (Kyrylkova et al., 2012). Further, a protein dye like Proteostat could be used to quantifiably examine protein aggregates in control compared to knockdown conditions (Navarro & Ventura, 2014). Developing planarians as a future model for studying neurodegeneration could prove pivotal in the field and would allow us to investigate neural loss in a system that is capable of robust regeneration to inform new approaches to tackling old questions.
Figure 3
Proteostasis Mechanisms for Neuronal Survival
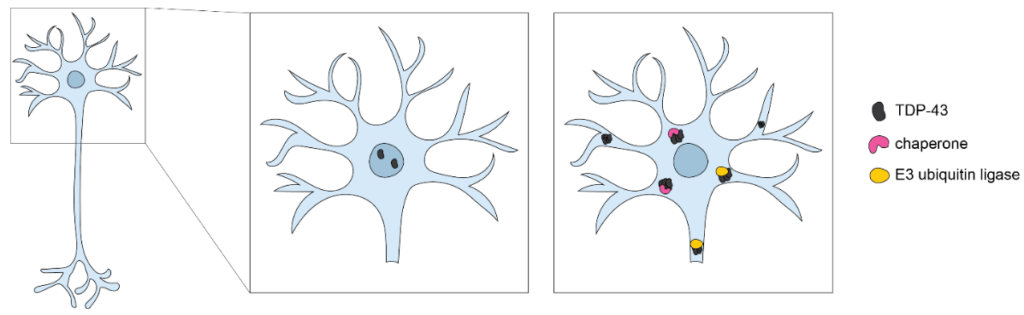
Note. Protein aggregates (black) accumulate in the cytoplasm, causing toxicity to neurons. Proteins involved in proteostasis (yellow and pink) work to prevent and break up protein aggregates.
Conclusion
Throughout the body of my thesis work, I uncovered molecular mechanisms of planarian nervous system regeneration and proposed the development of planarians as a future model for neurodegeneration. Specifically, I showed that combinatorial mechanisms specify neuronal neurotransmitter identity and cellular regionalization for planarian dopaminergic neurons. For the first time in planarians, I examined the cellular roles of the five planarian TDP-43 homologs and determined their importance in brain regeneration, stem cell maintenance, and function. Finally, I provided the starting point for a fascinating avenue to develop planarians as a model with which to study neurodegeneration in a physiologically relevant manner.
References
Agata, Kiyokazu; Soejima, Yukihiro; Kato, Kentaro; Kobayashi, Chiyoko; Umesono, Yoshihiko; & Watanabe, Kenji. (1998). Structure of the Planarian Central Nervous System (CNS) Revealed by Neuronal Cell Markers. Https://Doi.Org/10.2108/Zsj.15.433, 15(3), 433–440. https://doi.org/10.2108/ZSJ.15.433
Alberts, Bruce; Johnson, Alexander; Lewis, Julian; Raff, Martin; Roberts, Keith; & Walter, Peter. (2002). Neural Development. https://www.ncbi.nlm.nih.gov/books/NBK26814/
Alemany, Anna; Florescu, Maria; Baron, Chloé S.; Peterson-Maduro, Josi; & Van Oudenaarden, Alexander. (2018). Whole-organism clone tracing using single-cell sequencing. Nature, 556(7699), 108–112. https://doi.org/10.1038/nature25969
Amamoto, Ryoji; Huerta, Violeta Gisselle Lopez; Takahashi, Emi; Dai, Guangping; Grant, Aaron K.; Fu, Zhanyan; & Arlotta, Paola. (2016). Adult axolotls can regenerate original neuronal diversity in response to brain injury. ELife, 5(MAY2016). https://doi.org/10.7554/ELIFE.13998
Ament, Seth A.; Cortes-Gutierrez, Marcia; Herb, Brian R.; Mocci, Evelina; Colantuoni, Carlo; & McCarthy, Margaret M. (2023). A single-cell genomic atlas for maturation of the human cerebellum during early childhood. Science Translational Medicine, 15(721). https://doi.org/10.1126/SCITRANSLMED.ADE1283
Anderson, David J. (2001). Stem cells and pattern formation in the nervous system: The possible versus the actual. Neuron, 30(1), 19–35. https://doi.org/10.1016/S0896-6273(01)00260-4
Anderson, R. B.; Newgreen, D. F.; & Young, HM. (2013). Neural Crest and the Development of the Enteric Nervous System. https://www.ncbi.nlm.nih.gov/books/NBK6273/
Andersson, Elisabet; Tryggvason, Ulrika; Deng, Qiaolin; Friling, Stina; Alekseenko, Zhanna; Robert, Benoit; Perlmann, Thomas; & Ericson, Johan. (2006). Identification of Intrinsic Determinants of Midbrain Dopamine Neurons. Cell, 124(2), 393–405. https://doi.org/10.1016/J.CELL.2005.10.037
Arenas Gómez, Claudia M.; Sabin, Keith Z.; & Echeverri, Karen. (2020). Wound healing across the animal kingdom: Crosstalk between the immune system and the extracellular matrix. Developmental Dynamics, 249(7), 834. https://doi.org/10.1002/DVDY.178
Arendt, Detlev; Tosches, Maria Antonietta; & Marlow, Heather. (2016). From nerve net to nerve ring, nerve cord and brain-evolution of the nervous system. Nature Reviews Neuroscience, 17(1), 61–72. https://doi.org/10.1038/NRN.2015.15
Atabay, Kutay Deniz; LoCascio, Samuel A.; de Hoog, Thom; & Reddien, Peter W. (2018). Self-organization and progenitor targeting generate stable patterns in planarian regeneration. Science, 360(6387), 404–409. https://doi.org/10.1126/SCIENCE.AAP8179
Baguñ, Jaume. (2012). The planarian neoblast: the rambling history of its origin and some current black boxes. The International Journal of Developmental Biology, 56(1-2–3), 19–37. https://doi.org/10.1387/ijdb.113463jb
Baguñà, Jaume. (1976). Mitosis in the intact and regenerating planarian Dugesia mediterranea n.sp. II. Mitotic studies during regeneration, and a possible mechanism of blastema formation. Journal of Experimental Zoology, 195(1), 65–79. https://doi.org/10.1002/jez.1401950107
Baguñà, Jaume; & Ballester, Rafael. (1978). The nervous system in planarians: Peripheral and gastrodermal plexuses, pharynx innervation, and the relationship between central nervous system structure and the acoelomate organization. Journal of Morphology, 155(2), 237–252. https://doi.org/10.1002/JMOR.1051550208
Baguñà, Jaume; Saló, Emili; & Auladell, Carme. (1989). Regeneration and pattern formation in planarians. III. Evidence that neoblasts are totipotent stem cells and the source of blastema cells. Development, 107, 77–86.
Barker, Roger A.; Parmar, Malin; Studer, Lorenz; & Takahashi, Jun. (2017). Human Trials of Stem Cell-Derived Dopamine Neurons for Parkinson’s Disease: Dawn of a New Era. Cell Stem Cell, 21(5), 569–573. https://doi.org/10.1016/j.stem.2017.09.014
Barmaki, Haleh; Nourazarian, Alireza; & Khaki-Khatibi, Fatemeh. (2023). Proteostasis and neurodegeneration: a closer look at autophagy in Alzheimer’s disease. Frontiers in Aging Neuroscience, 15, 1281338. https://doi.org/10.3389/FNAGI.2023.1281338
Bazira, Peter J. (2021). An overview of the nervous system. Surgery (Oxford), 39(8), 451–462. https://doi.org/10.1016/J.MPSUR.2021.06.012
Bédard, Catherine; Wallman, Marie Josée; Pourcher, Emmanuelle; Gould, Peter V.; Parent, André; & Parent, Martin. (2011). Serotonin and dopamine striatal innervation in Parkinson’s disease and Huntington’s chorea. Parkinsonism and Related Disorders, 17(8), 593–598. https://doi.org/10.1016/j.parkreldis.2011.05.012
Benham-Pyle, Blair W.; Mann, Frederick G.; Brewster, Carolyn E.; Dewars, Enya R.; Vuu, Dung M.; Nowotarski, Stephanie H.; Guerrero-Hernández, Carlos; Malloy, Seth; Hall, Kate E.; Maddera, Lucinda E.; Chen, Shiyuan; Morrison, Jason A.; McKinney, Sean A.; Slaughter, Brian D.; Perera, Anoja; & Alvarado, Alejandro Sánchez. (2023). Planarians employ diverse and dynamic stem cell microenvironments to support whole-body regeneration. BioRxiv, 2022.03.20.485025. https://doi.org/10.1101/2022.03.20.485025
Bhat, Krishna Moorthi. (2007). Wingless activity in the precursor cells specifies neuronal migratory behavior in the Drosophila nerve cord. Developmental Biology, 311(2), 613–622. https://doi.org/10.1016/J.YDBIO.2007.09.004
Bisht, Kanchan; Sharma, Kaushik P.; Lecours, Cynthia; Gabriela Sánchez, Maria; El Hajj, Hassan; Milior, Giampaolo; Olmos-Alonso, Adrián; Gómez-Nicola, Diego; Luheshi, Giamal; Vallières, Luc; Branchi, Igor; Maggi, Laura; Limatola, Cristina; Butovsky, Oleg; & Tremblay, Marie Ève. (2016). Dark microglia: A new phenotype predominantly associated with pathological states. Glia, 64(5), 826–839. https://doi.org/10.1002/GLIA.22966
Björklund, A.; Dunnett, S. B.; Stenevi, U.; Lewis, M. E.; & Iversen, S. D. (1980). Reinnervation of the denervated striatum by substantia nigra transplants: Functional consequences as revealed by pharmacological and sensorimotor testing. Brain Research, 199(2), 307–333. https://doi.org/10.1016/0006-8993(80)90692-7
Björklund, A.; & Lindvall, O. (1984). Dopamine-containing systems in the CNS. Handbook of Chemical Neuroanatomy : Classical Transmitters in the CNS, 2, 55–122. https://cir.nii.ac.jp/crid/1571417125170103296
Björklund, A.; Stenevi, U.; Dunnett, S. B.; & Iversen, S. D. (1981). Functional reactivation of the deafferented neostriatum by nigral transplants. Nature, 289(5797), 497–499. https://doi.org/10.1038/289497A0
Björklund, Anders; & Stenevi, Ulf. (1979). Reconstruction of the nigrostriatal dopamine pathway by intracerebral nigral transplants. Brain Research, 177(3), 555–560. https://doi.org/10.1016/0006-8993(79)90472-4
Björklund, Anders; Stenevi, Ulf; & Svendgaard, Niels Aage. (1976). Growth of transplanted monoaminergic neurones into the adult hippocampus along the perforant path. Nature, 262(5571), 787–790. https://doi.org/10.1038/262787A0
Blackshaw, Seth; & Cayouette, Michel. (2025). Timing neural development and regeneration. Current Opinion in Neurobiology, 91, 102976. https://doi.org/10.1016/J.CONB.2025.102976
Blanchard, Veronique; Raisman‐Vozari, Rita; Savasta, Marc; Hirsch, Etienne; Javoy‐Agid, France; Feuerstein, Claude; & Agid, Yves. (1993). Cellular Quantification of Tyrosine Hydroxylase in the Rat Brain by Immunoautoradiography. Journal of Neurochemistry, 61(2), 617–626. https://doi.org/10.1111/J.1471-4159.1993.TB02166.X
Bonilla, Sonia; Hall, Anita C.; Pinto, Luisa; Attardo, Alessio; Götz, Magdalena; Huttner, Wieland B.; & Arenas, Ernest. (2008). Identification of midbrain floor plate radial glia-like cells as dopaminergic progenitors. GLIA, 56(8), 809–820. https://doi.org/10.1002/GLIA.20654
Bowling, Sarah; Sritharan, Duluxan; Osorio, Fernando G.; Nguyen, Maximilian; Cheung, Priscilla; Rodriguez-Fraticelli, Alejo; Patel, Sachin, Yuan; Wei Chien; Fujiwara, Yuko; Li, Bin E.; Orkin, Stuart H.; Hormoz, Sahand; & Camargo, Fernando D. (2020). An Engineered CRISPR-Cas9 Mouse Line for Simultaneous Readout of Lineage Histories and Gene Expression Profiles in Single Cells. Cell, 181(6), 1410-1422.e27. https://doi.org/10.1016/j.cell.2020.04.048
Braak, H.; Rüb, U.; Jansen Steur, E. N. H.; Del Tredici, K.; & De Vos, R. A. I. (2005). Cognitive status correlates with neuropathologic stage in Parkinson disease. Neurology, 64(8), 1404–1410. https://doi.org/10.1212/01.WNL.0000158422.41380.82
Braak, Heiko; & Del Tredici, Kelly. (2017). Neuropathological Staging of Brain Pathology in Sporadic Parkinson’s disease: Separating the Wheat from the Chaff. Journal of Parkinson’s Disease, 7(s1), S73–S87. https://doi.org/10.3233/JPD-179001
Braak, Heiko; Del Tredici, Kelly; Rüb, Udo; De Vos, Rob A. I.; Jansen Steur, Ernst N. H.; & Braak, Eva. (2003). Staging of brain pathology related to sporadic Parkinson’s disease. Neurobiology of Aging, 24(2), 197–211. https://doi.org/10.1016/S0197-4580(02)00065-9
Briscoe, James; Pierani, Alessandra; Jessell, Thomas M.; & Ericson, Johan. (2000). A homeodomain protein code specifies progenitor cell identity and neuronal fate in the ventral neural tube. Cell, 101(4), 435–445. https://doi.org/10.1016/S0092-8674(00)80853-3
Brundin, P.; Strecker, R. E.; Widner, H.; Clarke, D. J.; Nilsson, O. G.; Åstedt, B.; Lindvall, O.; & Björklund, A. (1988). Human fetal dopamine neurons grafted in a rat model of Parkinson’s disease: immunological aspects, spontaneous and drug-induced behaviour, and dopamine release. Experimental Brain Research, 70(1), 192–208. https://doi.org/10.1007/BF00271860
Burkhardt, Pawel. (2022). Ctenophores and the evolutionary origin(s) of neurons. Trends in Neurosciences, 45(12), 878–880. https://doi.org/10.1016/J.TINS.2022.09.001
Butt, Arthur; & Verkhratsky, Alexei. (2018). Neuroglia: Realising their true potential. Brain and Neuroscience Advances, 2. https://doi.org/10.1177/2398212818817495
Bylund, Magdalena; Andersson, Elisabeth; Novitch, Bennett G.; & Muhr, Jonas. (2003). Vertebrate neurogenesis is counteracted by Sox1-3 activity. Nature Neuroscience, 6(11), 1162–1168. https://doi.org/10.1038/NN1131
Caldwell, Lindsey J.; Davies, Nick O.; Cavone, Leonardo; Mysiak, Karolina S.; Semenova, Svetlana A.; Panula, Pertti; Armstrong, J. Douglas; Becker, Catherina G.; & Becker, Thomas. (2019). Regeneration of dopaminergic neurons in adult zebrafish depends on immune system activation and differs for distinct populations. Journal of Neuroscience, 39(24), 4694–4713. https://doi.org/10.1523/JNEUROSCI.2706-18.2019
Calne, D. B.; Teychenne, P. F.; Claveria, L. E.; Eastman, R.; Greenacre, J. K.; & Petrie, A. (1974). Bromocriptine in parkinsonism. British Medical Journal, 4(5942), 442–444. https://doi.org/10.1136/BMJ.4.5942.442
Carpenter, K. S.; Morita, M.; & Best, J. B. (1974). Ultrastructure of the photoreceptor of the planarian Dugesia dorotocephala – I. Normal eye. Cell and Tissue Research, 148(2), 143–158. https://doi.org/10.1007/BF00224579
Cascione, Mariafrancesca; De Matteis, Valeria;, Leporatti, Stefano; & Rinaldi, Rosaria. (2020). The new frontiers in neurodegenerative diseases treatment: Liposomal-based strategies. Frontiers in Bioengineering and Biotechnology, 8, 566767. 566767. https://doi.org/10.3389/fbioe.2020.566767
Cebrià, Francesc. (2016). Planarian body-wall muscle: Regeneration and function beyond a simple skeletal support. Frontiers in Cell and Developmental Biology, 4(FEB), 174749. https://doi.org/10.3389/fcell.2016.00008
Cebrià, Francesc; Guo, Tingxia; Jopek, Jessica; & Newmark, Phillip A. (2007). Regeneration and maintenance of the planarian midline is regulated by a slit orthologue. Developmental Biology, 307(2), 394. https://doi.org/10.1016/J.YDBIO.2007.05.006
Chalazonitis, Alcmène; Rao, Meenakshi; & Sulzer, David. (2022). Similarities and differences between nigral and enteric dopaminergic neurons unravel distinctive involvement in Parkinson’s disease. NPJ Parkinson’s Disease, 8(1), 50. https://doi.org/10.1038/S41531-022-00308-9
Chandra, Bidushi; Voas, Matthew G.; Davies, Erin L.; & Roberts-Galbraith, Rachel H. (2023). Ets-1 transcription factor regulates glial cell regeneration and function in planarians. Development (Cambridge), 150(18). https://doi.org/10.1242/dev.201666
Chhetri, Parvat Kuwar; & Das, Joe M. (2023). Neuroanatomy, Neural Tube Development and Stages. StatPearls. https://www.ncbi.nlm.nih.gov/books/NBK557414/
Chinta, Shankar J.; & Andersen, Julie K. (2005). Dopaminergic neurons. The International Journal of Biochemistry & Cell Biology, 37(5), 942–946. https://doi.org/10.1016/J.BIOCEL.2004.09.009
Cizeron, Mélissa; Qiu, Zhen; Koniaris, Babis; Gokhale, Ragini; Komiyama, Noboru H.; Fransén, Erik; & Grant, Seth G. N. (2020). A brainwide atlas of synapses across the mouse life span. Science, 369(6501). https://doi.org/10.1126/SCIENCE.ABA3163
Clay, Kendall B.; Medlock-Lanier, Taylor; Grimes, Rachel N.; Oke, Olabamibo O.; Filipov, Nikolay M.; & Roberts-Galbraith, Rachel H. (2025). Combinatorial mechanisms specify cellular location and neurotransmitter identity during regeneration of planarian neurons. BioRxiv, 2025.05.23.655781. https://doi.org/10.1101/2025.05.23.655781
Cobb, J. L. S. (1987). Neurobiology of the Echinodermata. In Nervous Systems in Invertebrates (pp. 483–525). Springer US. https://doi.org/10.1007/978-1-4613-1955-9_17
Colas, Jean François; & Schoenwolf, Gary C. (2001). Towards a cellular and molecular understanding of neurulation. Developmental Dynamics, 221(2), 117–145. https://doi.org/10.1002/DVDY.1144
Collins, James J.; Hou, Xiaowen; Romanova, Elena V.; Lambrus, Bramwell G.; Miller, Claire M.; Saberi, Amir; Sweedler, Jonathan V.; & Newmark, Phillip A. (2010). Genome-Wide Analyses Reveal a Role for Peptide Hormones in Planarian Germline Development. PLoS Biology, 8(10), e1000509. https://doi.org/10.1371/journal.pbio.1000509
Contreras, Estefania; Bolívar, Sara; Navarro, Xavier; & Udina, Esther. (2022). New insights into peripheral nerve regeneration: The role of secretomes. Experimental Neurology, 354, 114069. https://doi.org/10.1016/J.EXPNEUROL.2022.114069
Cotzias, George C.; Papavasiliou, Paul S.; Fehling, Clas,;Kaufman, Barry; & Mena, Ismael. (1970). Similarities between Neurologic Effects of L-Dopa and of Apomorphine. New England Journal of Medicine, 282(1), 31–33. https://doi.org/10.1056/NEJM197001012820107
Cousins, Fiona L.; Filby, Caitlin E.; & Gargett, Caroline E. (2022). Endometrial Stem/Progenitor Cells–Their Role in Endometrial Repair and Regeneration. Frontiers in Reproductive Health, 3, 811537. https://doi.org/10.3389/FRPH.2021.811537
Cowan, W. M. (1978). Aspects of neural development. International Review of Physiology, 17, 149–191.
Cowan, W. M. (1979). The development of the brain. Scientific American, 241(3), 113–133. https://doi.org/10.1038/SCIENTIFICAMERICAN0979-112
Cowles, Martis W.; Brown, David D. R.; Nisperos, Sean V.; Stanley, Brianna N.; Pearson, Bret J.; & Zayas, Ricardo M. (2013). Genome-wide analysis of the bHLH gene family in planarians identifies factors required for adult neurogenesis and neuronal regeneration. Development (Cambridge), 140(23), 4691–4702. https://doi.org/10.1242/DEV.098616
Cowles, Martis W.; Omuro, Kerilyn C.; Stanley, Brianna N.; Quintanilla, Carlo G.; & Zayas, Ricardo M. (2014). COE Loss-of-Function Analysis Reveals a Genetic Program Underlying Maintenance and Regeneration of the Nervous System in Planarians. PLoS Genetics, 10(10). https://doi.org/10.1371/journal.pgen.1004746
Currie, Ko W.; & Pearson, Bret J. (2013). Transcription factors lhx1/5-1 and pitx are required for the maintenance and regeneration of serotonergic neurons in planarians. Development, 140(17), 3577–3588. https://doi.org/10.1242/dev.098590
Damier, P.; Hirsch, E. C.,; Agid, Y.; & Graybiel, A. M. (1999). The substantia nigra of the human brain: II. Patterns of loss of dopamine-containing neurons in Parkinson’s disease. Brain, 122(8), 1437–1448. https://doi.org/10.1093/BRAIN/122.8.1437
Darras, Sébastien; Gerhart, John; Terasaki, Mark; Kirschner, Marc; & Lowe, Christopher J. (2011). β-Catenin specifies the endomesoderm and defines the posterior organizer of the hemichordate Saccoglossus kowalevskii. Development, 138(5), 959–970. https://doi.org/10.1242/DEV.059493
Dawson, Ted M.; Golde, Todd E.; & Lagier-Tourenne, Clotilde. (2018). Animal Models of Neurodegenerative Diseases. Nature Neuroscience, 21(10), 1370. https://doi.org/10.1038/S41593-018-0236-8
den Hartog Jager, W. A.; & Bethlem, J. (1960). The distribution of Lewy bodies in the central and autonomic nervous systems in idiopathic paralysis agitans. Journal of Neurology, Neurosurgery, and Psychiatry, 23(4), 283–290. https://doi.org/10.1136/JNNP.23.4.283
Development of the Central Nervous System – Spinal Cord – TeachMeAnatomy. (n.d.). Retrieved 28 March 2026, from https://teachmeanatomy.info/the-basics/embryology/central-nervous-system/
Díaz-Villanueva, José Fernando; Díaz-Molina, Raúl; & García-González, Victor. (2015). Protein Folding and Mechanisms of Proteostasis. International Journal of Molecular Sciences 2015, 16(8), 17193–17230. https://doi.org/10.3390/IJMS160817193
Doe, Chris Q. (2017). Temporal Patterning in the Drosophila CNS. Annual Review of Cell and Developmental Biology, 33, 219–240. https://doi.org/10.1146/ANNUREV-CELLBIO-111315-125210
Doitsidou, Maria; Flames, Nuria; Lee, Albert C.; Boyanov, Alexander; & Hobert, Oliver. (2008). Automated screening for mutants affecting dopaminergic-neuron specification in C. elegans. Nature Methods, 5(10), 869–872. https://doi.org/10.1038/nmeth.1250
Doitsidou, Maria; Flames, Nuria; Topalidou, Irini; Abe, Namiko; Felton, Terry; Remesal, Laura; Popovitchenko, Tatiana; Mann, Richard; Chalfie, Martin; & Hobert, Oliver. (2013). A combinatorial regulatory signature controls terminal differentiation of the dopaminergic nervous system in C. elegans. Genes & Development, 27(12), 1391–1405. https://doi.org/10.1101/gad.217224.113
Doitsidou, Maria; Minevich, Gregory; Kroll, Jason R.; Soete, Gwen; Gowtham, Sriharsh; Korswagen, Hendrik C.; van Zon, Jeroen Sebastiaan; & Hobert, Oliver. (2018). A Caenorhabditis elegans zinc finger transcription factor, ztf-6, required for the specification of a dopamine neuron-producing lineage. G3: Genes, Genomes, Genetics, 8(1), 17–26. https://doi.org/10.1534/G3.117.300132/-/DC1
Duman-Scheel, Molly; Li, Xuelin; Orlov, Irena; Noll, Markus; & Patel, Nipam H. (1997). Genetic separation of the neural and cuticular patterning functions of gooseberry. Development, 124(15), 2855–2865. https://doi.org/10.1242/DEV.124.15.2855
Durães, Fernando; Pinto, Madalena; & Sousa, Emília. (2018). Old Drugs as New Treatments for Neurodegenerative Diseases. Pharmaceuticals 2018, Vol. 11, Page 44, 11(2), 44. https://doi.org/10.3390/PH11020044
Duvoisin, Roger C. (1984). Is Parkinson’s Disease Acquired or Inherited? Canadian Journal of Neurological Sciences, 11(S1), 151–155. https://doi.org/10.1017/S031716710004631X
Eisenhoffer, George T.; Kang, Hara; & Alvarado, Alejandro Sánchez. (2008). Molecular Analysis of Stem Cells and Their Descendants during Cell Turnover and Regeneration in the Planarian Schmidtea mediterranea. Cell Stem Cell, 3(3), 327–339. https://doi.org/10.1016/J.STEM.2008.07.002
El-Danaf, Rana N.; Rajesh, Raghuvanshi; & Desplan, Claude. (2023). Temporal regulation of neural diversity in Drosophila and vertebrates. Seminars in Cell & Developmental Biology, 142, 13–22. https://doi.org/10.1016/J.SEMCDB.2022.05.011
Elkouris, Maximilianos; Balaskas, Nikos; Poulou, Maria; Politis, Panagiotis K.; Panayiotou, Elena; Malas, Stavros; Thomaidou, Dimitra; & Remboutsika, Eumorphia. (2011). Sox1 maintains the undifferentiated state of cortical neural progenitor cells via the suppression of Prox1-mediated cell cycle exit and neurogenesis. Stem Cells, 29(1), 89–98. https://doi.org/10.1002/STEM.554
Elliott, Jimmy; Jolicoeur, Christine; Ramamurthy, Vasanth; & Cayouette, Michel. (2008). Ikaros Confers Early Temporal Competence to Mouse Retinal Progenitor Cells. Neuron, 60(1), 26–39. https://doi.org/10.1016/J.NEURON.2008.08.008
Ellis, John. (1988). Proteins as molecular chaperones. Nature, 328(6129), 378–379.
Emili, Elena; Pallarès, Macià Esteve; Romero, Rafael; & Cebrià, Francesc. (2019). Smed-egfr-4 is required for planarian eye regeneration. International Journal of Developmental Biology, 63(1–2), 9–15. https://doi.org/10.1387/IJDB.180361FC
Fahn, Stanley. (2005). Does levodopa slow or hasten the rate of progression of Parkinson’s disease? Journal of Neurology, 252(SUPPL. 4). https://doi.org/10.1007/S00415-005-4008-5
Failli, Vieri; Bachy, Isabelle; & Rétaux, Sylvie. (2002). Expression of the LIM-homeodomain gene Lmx1a (dreher) during development of the mouse nervous system. Mechanisms of Development, 118(1–2), 225–228. https://doi.org/10.1016/S0925-4773(02)00254-X
Falck, B.; Hillarp, N. Å.; Thieme, G.; & Torp, A. (1962). Fluorescence of Catechol Amines and Related Compounds Condensed with Formaldehyde. Journal of Histochemistry & Cytochemistry, 10(3), 348–354. https://doi.org/10.1177/10.3.348
Fan, Guoping; & Katz, David M. (1993). Non-neuronal cells inhibit catecholaminergic differentiation of primary sensory neurons: role of leukemia inhibitory factor. Development, 118(1), 83–93. https://doi.org/10.1242/DEV.118.1.83
Faust, Travis E.; Gunner, Georgia; & Schafer, Dorothy P. (2021). Mechanisms governing activity-dependent synaptic pruning in the mammalian CNS. Nature Reviews. Neuroscience, 22(11), 657. https://doi.org/10.1038/S41583-021-00507-Y
Ferenczy, Alex; Bertrand, Gracia; & Gelfand, Morrie M. (1979). Proliferation kinetics of human endometrium during the normal menstrual cycle. American Journal of Obstetrics and Gynecology, 133(8), 859–867. https://doi.org/10.1016/0002-9378(79)90302-8
Fincher, Christopher T.; Wurtzel, Omri; de Hoog, Thom; Kravarik, Kellie M.; & Reddien, Peter W. (2018). Cell type transcriptome atlas for the planarian Schmidtea mediterranea. Science, 360(6391). https://doi.org/10.1126/science.aaq1736
Flames, Nuria; & Hobert, Oliver. (2009). Gene regulatory logic of dopamine neuron differentiation. Nature, 458(7240), 885–889. https://doi.org/10.1038/nature07929
Flores, Natasha M.; Oviedo, Néstor J.; & Sage, Julien. (2016). Essential role for the planarian intestinal GATA transcription factor in stem cells and regeneration. Developmental Biology, 418(1), 179. https://doi.org/10.1016/J.YDBIO.2016.08.015
Formery, Laurent; Orange, François; Formery, Antoine; Yaguchi, Shunsuke; Lowe, Christopher J.; Schubert, Michael; & Croce, Jenifer C. (2021). Neural anatomy of echinoid early juveniles and comparison of nervous system organization in echinoderms. The Journal of Comparative Neurology, 529(6), 1135–1156. https://doi.org/10.1002/CNE.25012
Forsthoefel, David J.; Cejda, Nicholas I.; Khan, Umair W.; & Newmark, Phillip A. (2020). Cell-type diversity and regionalized gene expression in the planarian intestine. ELife, 9. https://doi.org/10.7554/ELIFE.52613
Freed, Curt R.; Greene, Paul E.; Breeze, Robert E.; Tsai, Wei-Yann; DuMouchel, William; Kao, Richard; Dillon, Sandra; Winfield, Howard; Culver, Sharon; Trojanowski, John Q.; Eidelberg, David; & Fahn, Stanley. (2001). Transplantation of Embryonic Dopamine Neurons for Severe Parkinson’s Disease. New England Journal of Medicine, 344(10), 710–719. https://doi.org/10.1056/NEJM200103083441002
García-Arrarás, José E.; Rojas-Soto, Michelle; Jiménez, Luis B.; & Díaz-Miranda, Lucy. (2001). The enteric nervous system of echinoderms: unexpected complexity revealed by neurochemical analysis. The Journal of Experimental Biology, 204(Pt 5), 865–873. https://doi.org/10.1242/JEB.204.5.865
Gargett, Caroline E.; & Ye, Louie. (2012). Endometrial reconstruction from stem cells. Fertility and Sterility, 98(1), 11–20. https://doi.org/10.1016/J.FERTNSTERT.2012.05.004
Gaviño, Michael A.; & Reddien, Peter W. (2011). A Bmp/Admp regulatory circuit controls maintenance and regeneration of dorsal-ventral polarity in planarians. Current Biology, 21(4), 294–299. https://doi.org/10.1016/j.cub.2011.01.017
Ge, Mengmeng; Sheikhshahrokh, Amirhossein; Shi, Xiang; Zhang, Yu Hong; Xu, Zhiheng; & Wu, Qing Feng. (2022). A Spacetime Odyssey of Neural Progenitors to Generate Neuronal Diversity. Neuroscience Bulletin, 39(4), 645. https://doi.org/10.1007/S12264-022-00956-0
Gibbons, Christopher H. (2019). Basics of autonomic nervous system function. Handbook of Clinical Neurology, 160, 407–418. https://doi.org/10.1016/B978-0-444-64032-1.00027-8
Gilbert, Scott F. (2000a). An Introduction to Early Developmental Processes. https://www.ncbi.nlm.nih.gov/books/NBK9992/
Gilbert, Scott F. (2000b). Formation of the Neural Tube. https://www.ncbi.nlm.nih.gov/books/NBK10080/
Gilbert, Scott F. (2000c). Principles of Development: Life Cycles and Developmental Patterns. https://www.ncbi.nlm.nih.gov/books/NBK10125/
Glavic, Alvaro; Gómez-Skarmeta, José Luis; & Mayor, Roberto. (2002). The homeoprotein Xiro1 is required for midbrain-hindbrain boundary formation. Development, 129(7), 1609–1621. https://doi.org/10.1242/DEV.129.7.1609
Goldstein, David S.; & Holmes, Courtney. (2008). Neuronal Source of Plasma Dopamine. Clinical Chemistry, 54(11), 1864. https://doi.org/10.1373/CLINCHEM.2008.107193
Gómez-Skarmeta, José Luis; & Modolell, Juan. (2002). Iroquois genes: genomic organization and function in vertebrate neural development. Current Opinion in Genetics & Development, 12(4), 403–408. https://doi.org/10.1016/S0959-437X(02)00317-9
Gordon, Tessa. (2020). Peripheral Nerve Regeneration and Muscle Reinnervation. International Journal of Molecular Sciences, 21(22), 8652. https://doi.org/10.3390/IJMS21228652
Grandas, Francisco; Martínez-Martín, Pablo; & Linazasoro, Gurutz. (1998). Quality of life in patients with Parkinson’s disease who transfer from standard levodopa to sinemet CR: The STAR study. Journal of Neurology, Supplement, 245(1). https://doi.org/10.1007/PL00007736,
Grandel, Heiner; Kaslin, Jan; Ganz, Julia; Wenzel, Isabell; & Brand, Michael. (2006). Neural stem cells and neurogenesis in the adult zebrafish brain: Origin, proliferation dynamics, migration and cell fate. Developmental Biology, 295(1), 263–277. https://doi.org/10.1016/j.ydbio.2006.03.040
Grealish, Shane; Diguet, Elsa; Kirkeby, Agnete; Mattsson, Bengt; Heuer, Andreas; Bramoulle, Yann; Van Camp, Nadja; Perrier, Anselme L.; Hantraye, Philippe; Björklund, Anders; & Parmar, Malin. (2014). Human ESC-derived dopamine neurons show similar preclinical efficacy and potency to fetal neurons when grafted in a rat model of Parkinson’s disease. Cell Stem Cell, 15(5), 653–665. https://doi.org/10.1016/j.stem.2014.09.017
Grimmelikhuijzen, C. J.; & Westfall, J. A. (1995). The nervous systems of cnidarians. EXS, 72, 7–24. https://doi.org/10.1007/978-3-0348-9219-3_2
Grosskortenhaus, Ruth; Pearson, Bret J.; Marusich, Amanda; & Doe, Chris Q. (2005). Regulation of Temporal Identity Transitions in Drosophila Neuroblasts. Developmental Cell, 8(2), 193–202. https://doi.org/10.1016/J.DEVCEL.2004.11.019
Gurley, Kyle A.; Elliott, Sarah A.; Simakov, Oleg; Schmidt, Heiko A.; Holstein, Thomas W.; & Alvarado, Alejandro Sánchez. (2010). Expression of secreted Wnt pathway components reveals unexpected complexity of the planarian amputation response. Developmental Biology, 347(1), 24–39. https://doi.org/10.1016/J.YDBIO.2010.08.007
Gurley, Kyle A.; Rink, Jochen C.; & Alvarado, Alejandro Sánchez. (2008). β-catenin defines head versus tail identity during planarian regeneration and homeostasis. Science, 319(5861), 323–327. https://doi.org/10.1126/science.1150029
Hao, Marlene M.; Bornstein, Joel C.; Vanden Berghe, Pieter; Lomax, Alan E.; Young, Heather M.; & Foong, Jaime P. P. (2013). The emergence of neural activity and its role in the development of the enteric nervous system. Developmental Biology, 382(1), 365–374. https://doi.org/10.1016/J.YDBIO.2012.12.006
Hardin, Jeff; & King, Ryan S. (2008). The long and the short of Wnt signaling in C. elegans. Current Opinion in Genetics and Development, 18(4), 362–367. https://doi.org/10.1016/j.gde.2008.06.006
Hartenstein, Volker; Cruz, Louie; Lovick, Jennifer K.; & Guo, Ming. (2016). Developmental analysis of the dopamine-containing neurons of the Drosophila brain. The Journal of Comparative Neurology, 525(2), 363. https://doi.org/10.1002/CNE.24069
Hartenstein, Volker; & Wodarz, Andreas. (2013). Initial neurogenesis in Drosophila. Wiley Interdisciplinary Reviews. Developmental Biology, 2(5), 701–721. https://doi.org/10.1002/WDEV.111
Hartl, F. U.; Martin, J.; & Neupert, W. (1992). Protein Folding in the Cell. Annual Review of Biophysics and Biomolecular Structure, 21, 293–322. https://doi.org/10.1146/ANNUREV.BB.21.060192.001453
Hartl, F. Ulrich. (1996). Molecular chaperones in cellular protein folding. Nature, 381(6583), 571–580. https://doi.org/10.1038/381571A0
Haugh, Bruce N. (1975). Nervous systems of Mississippian camerate crinoids. Paleobiology, 1(3), 261–272. https://doi.org/10.1017/S0094837300002529
Hayashi, Tetsutaro; Asami, Maki; Higuchi, Sayaka; Shibata, Norito; & Agata, Kiyokazu. (2006). Isolation of planarian X-ray-sensitive stem cells by fluorescence-activated cell sorting. Development, Growth and Differentiation, 48(6), 371–380. https://doi.org/10.1111/j.1440-169X.2006.00876.x
He, Lingjuan; Pu, Wenjuan; Liu, Xiuxiu; Zhang, Zhenqian; Han, Maoying; Li, Yi; Huang, Xiuzhen; Han, Ximeng; Li, Yan; Liu, Kuo; Shi, Mengyang; Lai, Liang; Sun, Ruilin; Wang, Qing Dong; Ji, Yong; Tchorz, Jan S.; & Zhou, Bin. (2021). Proliferation tracing reveals regional hepatocyte generation in liver homeostasis and repair. Science, 371(6532). https://doi.org/10.1126/science.abc4346
Hechst, B.; & Nussbaum, L. (1931). Beiträge zur Histopathologie der sympathischen Ganglien. Archiv Für Psychiatrie Und Nervenkrankheiten, 95(1), 556–583. https://doi.org/10.1007/BF01819126
Hirsch, Etienne; Graybiel, Ann M.; & Agid, Yves A. (1988). Melanized dopaminergic neurons are differentially susceptible to degeneration in Parkinson’s disease. Nature, 334(6180), 345–348. https://doi.org/10.1038/334345a0
Holguera, Isabel; & Desplan, Claude. (2018). Neuronal specification in space and time. Science, 362(6411), 176–180. https://doi.org/10.1126/SCIENCE.AAS9435
Huang, Carol; Chan, Jennifer A.; & Schuurmans, Carol. (2014). Proneural bHLH Genes in Development and Disease. Current Topics in Developmental Biology, 110, 75–127. https://doi.org/10.1016/B978-0-12-405943-6.00002-6
Hynes, Mary; & Rosenthal, Arnon. (1999). Specification of dopaminergic and serotonergic neurons in the vertebrate CNS. Current Opinion in Neurobiology, 9(1), 26–36. https://doi.org/10.1016/S0959-4388(99)80004-X
Iglesias, Marta; Almuedo-Castillo, Maria; Aboobaker, A. Aziz; & Saló, Emili. (2011). Early planarian brain regeneration is independent of blastema polarity mediated by the Wnt/β-catenin pathway. Developmental Biology, 358(1), 68–78. https://doi.org/10.1016/j.ydbio.2011.07.013
Iglesias, Marta; Gomez-Skarmeta, Jose Luis; Saló, Emili; & Adell, Teresa. (2008). Silencing of Smed-βcatenin1 generates radial-like hypercephalized planarians. Development, 135(7), 1215–1221. https://doi.org/10.1242/DEV.020289
Iismaa, Siiri E.; Kaidonis, Xenia; Nicks, Amy M.; Bogush, Nikolay; Kikuchi, Kazu; Naqvi, Nawazish; Harvey, Richard P.; Husain, Ahsan; & Graham, Robert M. (2018). Comparative regenerative mechanisms across different mammalian tissues. Npj Regenerative Medicine, 3(1), 1–20. https://doi.org/10.1038/s41536-018-0044-5
Inoue, Takeshi; Kumamoto, Hiroshi; Okamoto, Keiji; Umesono, Yoshihiko; Sakai, Masaki; Alvarado, Alejandro Sánchez; & Agata, Kiyokazu. (2004). Morphological and Functional Recovery of the Planarian Photosensing System during Head Regeneration. Zoological Science, 21(3), 275–283. https://doi.org/10.2108/zsj.21.275
Iranzo, Alex; Gelpi, Ellen; Tolosa, Eduard; Molinuevo, José Luis; Serradell, Mónica; Gaig, Carles; & Santamaria, Joan. (2014). Neuropathology of prodromal Lewy body disease. Movement Disorders, 29(3), 410–415. https://doi.org/10.1002/MDS.25825
Isshiki, Takako; Pearson, Bret; Holbrook, Scott; & Doe, Chris Q. (2001). Drosophila Neuroblasts Sequentially Express Transcription Factors which Specify the Temporal Identity of Their Neuronal Progeny. Cell, 106(4), 511–521. https://doi.org/10.1016/S0092-8674(01)00465-2
Itoh, Motoyuki; Kudoh, Tetsuhiro; Dedekian, Michael; Kim, Cheol Hee; & Chitnis, Ajay B. (2002). A role for iro1 and iro7 in the establishment of an anteroposterior compartment of the ectoderm adjacent to the midbrain-hindbrain boundary. Development, 129(10), 2317–2327. https://doi.org/10.1242/DEV.129.10.2317
Javed, Awais; Mattar, Pierre; Lu, Suying; Kruczek, Kamil; Kloc, Magdalena; Gonzalez-Cordero, Anai; Bremner, Rod; Ali, Robin R.,; & Cayouette, Michel. (2020). Pou2f1 and Pou2f2 cooperate to control the timing of cone photoreceptor production in the developing mouse retina. Development (Cambridge, England), 147(18). https://doi.org/10.1242/DEV.188730
Javed, Awais; Santos-França, Pedro L.; Mattar, Pierre; Cui, Allie; Kassem, Fatima; & Cayouette, Michel. (2023). Ikaros family proteins redundantly regulate temporal patterning in the developing mouse retina. Development (Cambridge, England), 150(2). https://doi.org/10.1242/DEV.200436
Joksimovic, Milan; Yun, Beth A.; Kittappa, Raja; Anderegg, Angela M.; Chang, Wendy W.; Taketo, Makoto M.; McKay, Ronald D. G.; & Awatramani, Rajeshwar B. (2009). Wnt antagonism of Shh facilitates midbrain floor plate neurogenesis. Nature Neuroscience, 12(2), 125–131. https://doi.org/10.1038/NN.2243
Jones, D.; Lowe, V.; Graff-Radford, J.; Botha, H.; Barnard, L.; Wiepert, D.; Murphy, M. C.; Murray, M.; Senjem, M.; Gunter, J.; Wiste, H.; Boeve, B.; Knopman, D.; Petersen, R.; & Jack, C. (2022). A computational model of neurodegeneration in Alzheimer’s disease. Nature Communications, 13(1), 1–13. https://doi.org/10.1038/s41467-022-29047-4
Ju, Dr William. (2020). 1.4 The Somatic Nervous System. https://pressbooks.claremont.edu/neurosciencecdn2/chapter/anatomy-physiology-the-somatic-nervous-system/
Kageyama, Ryoichiro; Ohtsuka, Toshiyuki; Shimojo, Hiromi; & Imayoshi, Itaru. (2009). Dynamic regulation of Notch signaling in neural progenitor cells. Current Opinion in Cell Biology, 21(6), 733–740. https://doi.org/10.1016/j.ceb.2009.08.009
Kameda, Yoko; Saitoh, Takayoshi; & Fujimura, Takao. (2011). Hes1 regulates the number and anterior–posterior patterning of mesencephalic dopaminergic neurons at the mid/hindbrain boundary (isthmus). Developmental Biology, 358(1), 91–101. https://doi.org/10.1016/J.YDBIO.2011.07.016
Katz, D. M.; Markey, K. A.; Goldstein, M.; & Black, I. B. (1983). Expression of catecholaminergic characteristics by primary sensory neurons in the normal adult rat in vivo. Proceedings of the National Academy of Sciences of the United States of America, 80(11 I), 3526–3530. https://doi.org/10.1073/PNAS.80.11.3526
Khakh, Baljit S.; & Deneen, Benjamin. (2019). The Emerging Nature of Astrocyte Diversity. Annual Review of Neuroscience, 42(Volume 42, 2019), 187–207. https://doi.org/10.1146/annurev-neuro-070918-050443
Kim, Hyun Jung; Sugimori, Michiya; Nakafuku, Masato; & Svendsen, Clive N. (2007). Control of neurogenesis and tyrosine hydroxylase expression in neural progenitor cells through bHLH proteins and Nurr1. Experimental Neurology, 203(2), 394–405. https://doi.org/10.1016/J.EXPNEUROL.2006.08.029
Kim, Jaesang; Lo, Liching; Dormand, Emma; & Anderson, David J. (2003). SOX10 Maintains Multipotency and Inhibits Neuronal Differentiation of Neural Crest Stem Cells. Neuron, 38(1), 17–31. https://doi.org/10.1016/S0896-6273(03)00163-6
Kim, Jong Hoon; Auerbach, Jonathan M.; Rodríguez-Gómez, José A.; Velasco, Iván; Gavin, Denise; Lumelsky, Nadya; Lee, Sang Hun; Nguyen, John; Sánchez-Pernaute, Rosario; Bankiewicz, Krys; & McKay, Ron. (2002). Dopamine neurons derived from embryonic stem cells function in an animal model of Parkinson’s disease. Nature, 418(6893), 50–56. https://doi.org/10.1038/NATURE00900,
King, Hunter O.; Owusu-Boaitey, Kwadwo E.; Fincher, Christopher T.; & Reddien, Peter W. (2024). A transcription factor atlas of stem cell fate in planarians. Cell Reports, 43(3). https://doi.org/10.1016/j.celrep.2024.113843
Kirkeby, Agnete; Grealish, Shane; Wolf, Daniel A.; Nelander, Jenny; Wood, James; Lundblad, Martin; Lindvall, Olle; & Parmar, Malin. (2012). Generation of Regionally Specified Neural Progenitors and Functional Neurons from Human Embryonic Stem Cells under Defined Conditions. Cell Reports, 1(6), 703–714. https://doi.org/10.1016/j.celrep.2012.04.009
Klein, Marianne O.; Battagello, Daniella S.; Cardoso, Ariel R.; Hauser, David N.; Bittencourt, Jackson C.; & Correa, Ricardo G. (2018). Dopamine: Functions, Signaling, and Association with Neurological Diseases. Cellular and Molecular Neurobiology, 39(1), 31. https://doi.org/10.1007/S10571-018-0632-3
Kordower, Jeffrey H.; Chu, Yaping; Hauser, Robert A.; Freeman, Thomas B.; & Olanow, C. Warren. (2008). Lewy body-like pathology in long-term embryonic nigral transplants in Parkinson’s disease. Nature Medicine, 14(5), 504–506. https://doi.org/10.1038/nm1747
Koszła, Oliwia; & Sołek, Przemysław. (2024). Misfolding and aggregation in neurodegenerative diseases: protein quality control machinery as potential therapeutic clearance pathways. Cell Communication and Signaling 2024 22:1, 22(1), 1–23. https://doi.org/10.1186/S12964-024-01791-8
Kraemer, Brian C.; Schuck, Theresa; Wheeler, Jeanna M.; Robinson, Linda C.; Trojanowski, John Q.; Lee, Virginia M. Y.; & Schellenberg, Gerard D. (2010). Loss of murine TDP-43 disrupts motor function and plays an essential role in embryogenesis. Acta Neuropathologica, 119(4), 409. https://doi.org/10.1007/S00401-010-0659-0
Kriks, Sonja; Shim, Jae Won; Piao, Jinghua; Ganat, Yosif M.; Wakeman, Dustin R.; Xie, Zhong; Carrillo-Reid, Luis; Auyeung, Gordon; Antonacci, Chris; Buch, Amanda; Yang, Lichuan, Beal, M. Flint; Surmeier, D. James; Kordower, Jeffrey H.; Tabar, Viviane; & Studer, Lorenz. (2011). Dopamine neurons derived from human ES cells efficiently engraft in animal models of Parkinson’s disease. Nature, 480(7378), 547–551. https://doi.org/10.1038/NATURE10648
Krugelis MacRae, Edith. (1964). Observations on the fine structure of photoreceptor cells in the planarian Dugesia tigrina. Journal of Ultrastructure Research, 10(3–4), 334–349. https://doi.org/10.1016/S0022-5320(64)80013-7
Kupsky, W. J.; Grimes, M. M.; Sweeting, J.; Bertsch, R.; & Cote, L. J. (1987). Parkinson’s disease and megacolon: Concentric hyaline inclusions (lewy bodies) in enteric ganglion cells. Neurology, 37(7), 1253–1255. https://doi.org/10.1212/wnl.37.7.1253
Kyrylkova, Kateryna; Kyryachenko, Sergiy; Leid, Mark; & Kioussi, Chrissa. (2012). Detection of Apoptosis by TUNEL Assay. Methods in Molecular Biology, 887, 41–47. https://doi.org/10.1007/978-1-61779-860-3_5
Lapan, Sylvain W.; & Reddien, Peter W. (2011). dlx and sp6-9 Control Optic Cup Regeneration in a Prototypic Eye. PLoS Genetics, 7(8), e1002226. https://doi.org/10.1371/journal.pgen.1002226
Lapan, Sylvain W.; & Reddien, Peter W. (2012a). Transcriptome analysis of the planarian eye identifies ovo as a specific regulator of eye regeneration. Cell Reports, 2(2), 294. https://doi.org/10.1016/J.CELREP.2012.06.018
Lapan, Sylvain W.; & Reddien, Peter W. (2012b). Transcriptome Analysis of the Planarian Eye Identifies ovo as a Specific Regulator of Eye Regeneration. Cell Reports, 2(2), 294–307. https://doi.org/10.1016/j.celrep.2012.06.018
Law, Simon W.; Conneely, Orla M.; DeMayo, Franco J.; & O’Malley, Bert W. (1992). Identification of a new brain-specific transcription factor, NURR1. Molecular Endocrinology, 6(12), 2129–2135. https://doi.org/10.1210/MEND.6.12.1491694,
Le Douarin, N. M.; & Teillet, M. A. (1973). The migration of neural crest cells to the wall of the digestive tract in avian embryo. Development, 30(1), 31–48. https://doi.org/10.1242/DEV.30.1.31
Le Douarin, Nicole M.; & Teillet, Marie Aimée M. (1974). Experimental analysis of the migration and differentiation of neuroblasts of the autonomic nervous system and of neurectodermal mesenchymal derivatives, using a biological cell marking technique. Developmental Biology, 41(1), 162–184. https://doi.org/10.1016/0012-1606(74)90291-7
Lecaudey, Virginie; Anselme, Isabelle; Rosa, Frédéric; & Schneider-Maunoury, Sylvie. (2004). The zebrafish Iroquois gene iro7 positions the r4/r5 boundary and controls neurogenesis in the rostral hindbrain. Development (Cambridge, England), 131(13), 3121–3131. https://doi.org/10.1242/DEV.01190
Lesage, Suzanne; & Brice, Alexis. (2009). Parkinson’s disease: from monogenic forms to genetic susceptibility factors. Human Molecular Genetics, 18(R1), R48–R59. https://doi.org/10.1093/HMG/DDP012
Li, Jia Yi; Englund, Elisabet; Holton, Janice L.; Soulet, Denis, Hagell, Peter; Lees, Andrew J.; Lashley, Tammaryn; Quinn, Niall P.; Rehncrona, Stig; Björklund, Anders; Widner, Håkan; Revesz, Tamas; Lindvall, Olle; & Brundin, Patrik. (2008). Lewy bodies in grafted neurons in subjects with Parkinson’s disease suggest host-to-graft disease propagation. Nature Medicine, 14(5), 501–503. https://doi.org/10.1038/NM1746
Li, X.; & Noll, M. (1993). Role of the gooseberry gene in Drosophila embryos: maintenance of wingless expression by a wingless–gooseberry autoregulatory loop. The EMBO Journal, 12(12), 4499–4509. https://doi.org/10.1002/j.1460-2075.1993.tb06139.x
Li, Z. S.; Pham, T. D.; Tamir, H.; Chen, J. J.; & Gershon, M. D. (2004). Enteric Dopaminergic Neurons: Definition, Developmental Lineage, and Effects of Extrinsic Denervation. The Journal of Neuroscience, 24(6), 1330. https://doi.org/10.1523/JNEUROSCI.3982-03.2004
Lodato, Simona; & Arlotta, Paola. (2015). Generating Neuronal Diversity in the Mammalian Cerebral Cortex. Annual Review of Cell and Developmental Biology, 31(Volume 31, 2015), 699–720. https://doi.org/10.1146/annurev-cellbio-100814-125353
Lodato, Simona; Rouaux, Caroline; Quast, Kathleen B.; Jantrachotechatchawan, Chanati; Studer, Michèle; Hensch, Takao K.; & Arlotta, Paola. (2011). Excitatory Projection Neuron Subtypes Control the Distribution of Local Inhibitory Interneurons in the Cerebral Cortex. Neuron, 69(4), 763–779. https://doi.org/10.1016/j.neuron.2011.01.015
Logan, Catriona Y.; & Nusse, Roel. (2004). The Wnt signaling pathway in development and disease. Annual Review of Cell and Developmental Biology, 20(Volume 20, 2004), 781–810. https://doi.org/10.1146/annurev.cellbio.20.010403.113126
Lucila Scimone, M.; Cote, Lauren E.; & Reddien, Peter W. (2017). Orthogonal muscle fibres have different instructive roles in planarian regeneration. Nature, 551(7682), 623–628. https://doi.org/10.1038/NATURE24660
Luo, Wenjie; Sun, Weilin; Taldone, Tony; Rodina, Anna; & Chiosis, Gabriela. (2010). Heat shock protein 90 in neurodegenerative diseases. Molecular Neurodegeneration, 5(1), 24. https://doi.org/10.1186/1750-1326-5-24
MacRae, Edith Krugelis. (1967). The fine structure of sensory receptor processes in the auricular epithelium of the planarian, Dugesia tigrina. Zeitschrift Für Zellforschung Und Mikroskopische Anatomie, 82(4), 479–494. https://doi.org/10.1007/BF00337119/METRICS
Maloney, Michael T.; & Bamburg, James R. (2011). Mechanisms of neuronal growth cone guidance: an historical perspective. Developmental Neurobiology, 71(9), 795. https://doi.org/10.1002/DNEU.20908
Marcos, C.; & Pachnis, V. (1996). The effect of the ret- mutation on the normal development of the central and parasympathetic nervous systems. The International Journal of Developmental Biology, Suppl 1, 137S-138S.
Marikawa, Yusuke. (2006). Wnt/β-catenin signaling and body plan formation in mouse embryos. Seminars in Cell and Developmental Biology, 17(2), 175–184. https://doi.org/10.1016/j.semcdb.2006.04.003
Marlow, Heather; Matus, David Q.; & Martindale, Mark Q. (2013). Ectopic activation of the canonical wnt signaling pathway affects ectodermal patterning along the primary axis during larval development in the anthozoan Nematostella vectensis. Developmental Biology, 380(2), 324–334. https://doi.org/10.1016/j.ydbio.2013.05.022
März, Martin; Seebeck, Florian; & Bartscherer, Kerstin. (2013). A pitx transcription factor controls the establishment and maintenance of the serotonergic lineage in planarians. Development (Cambridge), 140(22), 4499–4509. https://doi.org/10.1242/DEV.100081
Matsumine, Hiroto; Yamamura, Yasuhiro; Hattori, Nobutaka; Kobayashi, Tomonori; Kitada, Tohru; Yoritaka, Asako; & Mizuno, Yoshikuni. (1998). A microdeletion of D6S305 in a family of autosomal recessive juvenile parkinsonism (PARK2). Genomics, 49(1), 143–146. https://doi.org/10.1006/geno.1997.5196
Mayor, Roberto; & Theveneau, Eric. (2013). The neural crest. Development, 140(11), 2247–2251. https://doi.org/10.1242/DEV.091751
McKenna, Aaron; Findlay, Gregory M.; Gagnon, James A.; Horwitz, Marshall S.; Schier, Alexander F.; & Shendure, Jay. (2016). Whole-organism lineage tracing by combinatorial and cumulative genome editing. Science, 353(6298). https://doi.org/10.1126/science.aaf7907
Medlock-Lanier, Taylor; Clay, Kendall B.; & Roberts-Galbraith, Rachel H. (2024). Planarian LDB and SSDP proteins scaffold transcriptional complexes for regeneration and patterning. Developmental Biology, 515, 67. https://doi.org/10.1016/J.YDBIO.2024.06.021
Melamed, Eldad. (1979). Early-Morning Dystonia: A Late Side Effect of Long-term Levodopa Therapy in Parkinson’s Disease. Archives of Neurology, 36(5), 308–310. https://doi.org/10.1001/ARCHNEUR.1979.00500410086014
Memic, Fatima; Knoflach, Viktoria; Sadler, Rebecca; Tegerstedt, Gunilla; Sundström, Erik; Guillemot, Francois; Pachnis, Vassilis; & Marklund, Ulrika. (2016). Ascl1 Is Required for the Development of Specific Neuronal Subtypes in the Enteric Nervous System. The Journal of Neuroscience, 36(15), 4339. https://doi.org/10.1523/JNEUROSCI.0202-16.2016
Mercurio, Silvia; Gattoni, Giacomo; Scarì, Giorgio; Ascagni, Miriam; Barzaghi, Benedetta; Elphick, Maurice R.; Croce, Jenifer C.; Schubert, Michael; Benito-Gutiérrez, Elia; & Pennati, Roberta. (2024). A feather star is born: embryonic development and nervous system organization in the crinoid Antedon mediterranea. Open Biology, 14(8), 240115. https://doi.org/10.1098/RSOB.240115
Molina, M. Dolores; Abduljabbar, Dema; Guixeras, Anna; Fraguas, Susanna; & Cebrià, Francesc. (2023). LIM-HD transcription factors control axial patterning and specify distinct neuronal and intestinal cell identities in planarians. Open Biology, 13(12), 230327. https://doi.org/10.1098/RSOB.230327
Molina, Ma Dolores; Saló, Emili; & Cebrià, Francesc. (2007). The BMP pathway is essential for re-specification and maintenance of the dorsoventral axis in regenerating and intact planarians. Developmental Biology, 311(1), 79–94. https://doi.org/10.1016/j.ydbio.2007.08.019
Molinaro, Alyssa M.; & Pearson, Bret J. (2016). In silico lineage tracing through single cell transcriptomics identifies a neural stem cell population in planarians. Genome Biology, 17(1), 1–17. https://doi.org/10.1186/s13059-016-0937-9
Moore, R. Y.; & Bloom, F. E. (1979). Central catecholamine neuron systems: anatomy and physiology of the norepinephrine and epinephrine systems. Annual Review of Neuroscience, 2(Volume 2, 1979), 113–168. https://doi.org/10.1146/annurev.ne.02.030179.000553
Moriarty, Niamh; Gantner, Carlos W.; Hunt, Cameron P. J.; Ermine, Charlotte M.; Frausin, Stefano; Viventi, Serena; Ovchinnikov, Dmitry A.; Kirik, Deniz; Parish, Clare L.; & Thompson, Lachlan H. (2022). A combined cell and gene therapy approach for homotopic reconstruction of midbrain dopamine pathways using human pluripotent stem cells. Cell Stem Cell, 29(3), 434-448.e5. https://doi.org/10.1016/j.stem.2022.01.013
Morita, M.; Best, J. B.; & Noel, J. (1969). Electron microscopic studies of planarian regeneration. I. Fine structure of neoblasts in Dugesia dorotocephala. Journal of Ultrastructure Research, 27(1), 7–23.
Muhr, Jeremy; Arbor, Tafline C.; & Ackerman, Kristin M. (2023). Embryology, Gastrulation. StatPearls. https://www.ncbi.nlm.nih.gov/books/NBK554394/
Nagarajan, Archana; Ning, Ye; Reisner, Kaja; Buraei, Zafir; Larsen, Jan Petter; Hobert, Oliver; & Doitsidou, Maria. (2014). Progressive degeneration of dopaminergic neurons through TRP channel-induced cell death. Journal of Neuroscience, 34(17), 5738–5746. https://doi.org/10.1523/JNEUROSCI.4540-13.2014
Nambu, John R.; Franks, Robert G.; Hu, Song; & Crews, Stephen T. (1990). The single-minded gene of Drosophila is required for the expression of genes important for the development of CNS midline cells. Cell, 63(1), 63–75. https://doi.org/10.1016/0092-8674(90)90288-P
Navarro, Susanna; & Ventura, Salvador. (2014). Fluorescent dye ProteoStat to detect and discriminate intracellular amyloid-like aggregates in Escherichia coli. Biotechnology Journal, 9(10), 1259–1266. https://doi.org/10.1002/BIOT.201400291
Niehrs, Christof. (2010). On growth and form: A Cartesian coordinate system of Wnt and BMP signaling specifies bilaterian body axes. Development, 137(6), 845–857. https://doi.org/10.1242/DEV.039651
Nishimura, K.; Kitamura, Y.; Taniguchi, T.; & Agata, K. (2010). Analysis of motor function modulated by cholinergic neurons in planarian Dugesia japonica. Neuroscience, 168(1), 18–30. https://doi.org/10.1016/j.neuroscience.2010.03.038
Nishimura, K.; Kitamura, Y.; Umesono, Y.; Takeuchi, K.; Takata, K.; Taniguchi, T.; & Agata, K. (2008). Identification of glutamic acid decarboxylase gene and distribution of GABAergic nervous system in the planarian Dugesia japonica. Neuroscience, 153(4), 1103–1114. https://doi.org/10.1016/J.NEUROSCIENCE.2008.03.026
Nishimura, Kaneyasu; Kitamura, Yoshihisa; Inoue, Takeshi; Umesono, Yoshihiko; Sano, Shozo; Yoshimoto, Kanji; Inden, Masatoshi; Takata, Kazuyuki; Taniguchi, Takashi; Shimohama, Shun; & Agata, Kiyokazu. (2007). Reconstruction of dopaminergic neural network and locomotion function in planarian regenerates. Developmental Neurobiology, 67(8), 1059–1078. https://doi.org/10.1002/dneu.20377
Nishimura, Kaneyasu; Kitamura, Yoshihisa; Inoue, Takeshi; Umesono, Yoshihiko; Yoshimoto, Kanji; Takeuchi, Kosei; Taniguchi, Takashi; & Agata, Kiyokazu. (2007). Identification and distribution of tryptophan hydroxylase (TPH)-positive neurons in the planarian Dugesia japonica. Neuroscience Research, 59(1), 101–106. https://doi.org/10.1016/J.NEURES.2007.05.014
Nishimura, Kaneyasu; Kitamura, Yoshihisa; Inoue, Takeshi; Umesono, Yoshihiko; Yoshimoto, Kanji; Taniguchi, Takashi; & Agata, Kiyokazu. (2008). Characterization of tyramine β-hydroxylase in planarian Dugesia japonica: Cloning and expression. Neurochemistry International, 53(6–8), 184–192. https://doi.org/10.1016/j.neuint.2008.09.006
Nobili, Annalisa; Latagliata, Emanuele Claudio; Viscomi, Maria Teresa; Cavallucci, Virve; Cutuli, Debora; Giacovazzo, Giacomo; Krashia, Paraskevi; Rizzo, Francesca Romana; Marino, Ramona; Federici, Mauro; De Bartolo, Paola; Aversa, Daniela; Dell’Acqua, Maria Concetta; Cordella, Alberto; Sancandi, Marco; Keller, Flavio; Petrosini, Laura; Puglisi-Allegra, Stefano; Mercuri, Nicola Biagio; … D’Amelio, Marcello. (2017). Dopamine neuronal loss contributes to memory and reward dysfunction in a model of Alzheimer’s disease. Nature Communications 2017 8:1, 8(1), 1–14. https://doi.org/10.1038/ncomms14727
Oberheim, Nancy Ann; Goldman, Steven A.; & Nedergaard, Maiken. (2012). Heterogeneity of astrocytic form and function. Methods in Molecular Biology, 814, 23–45. https://doi.org/10.1007/978-1-61779-452-0_3
Ogawa, Kazuya; Ishihara, Shogo; Saito, Yumi; Mineta, Katsuhiko; Nakazawa, Masum;, Ikeo, Kazuho; Gojobori, Takashi; Watanabe, Kenji; & Agata, Kiyokazu. (2002). Induction of a noggin-Like Gene by Ectopic DV Interaction during Planarian Regeneration. Developmental Biology, 250(1), 59–70. https://doi.org/10.1006/DBIO.2002.0790
Olanow, C. Warren; Goetz, Christopher G.; Kordower, Jeffrey H.; Stoessl, A. Jon; Sossi, Vesna; Brin, Mitchell F.; Shannon, Kathleen M.; Nauert, G. Michael; Perl, Daniel P.; Godbold, James; & Freeman, Thomas B. (2003). A double-blind controlled trial of bilateral fetal nigral transplantation in Parkinson’s disease. Annals of Neurology, 54(3), 403–414. https://doi.org/10.1002/ANA.10720,
Ong, Ta Hsuan; Romanova, Elena V.; Roberts-Galbraith, Rachel H.; Yang, Ning; Zimmerman, Tyler A.; Collins, James J.; Lee, Ji Eun; Kelleher, Neil L.; Newmark, Phillip A.; & Sweedler, Jonathan V. (2016). Mass spectrometry imaging and identification of peptides associated with cephalic ganglia regeneration in Schmidtea mediterranea. Journal of Biological Chemistry, 291(15), 8109–8120. https://doi.org/10.1074/jbc.M115.709196
Ono, Yuichi; Nakatani, Tomoya; Minaki, Yasuko; & Kumai, Minoru. (2010). The basic helix-loop-helix transcription factor Nato3 controls neurogenic activity in mesencephalic floor plate cells. Development, 137(11), 1897–1906. https://doi.org/10.1242/DEV.042572
Ono, Yuichi; Nakatani, Tomoya; Sakamoto, Yoshimasa; Mizuhara, Eri; Minaki, Yasuko; Kumai, Minoru; Hamaguchi, Akiko; Nishimura, Miyuki; Inoue, Yoko; Hayashi, Hideki; Takahashi, Jun; & Imai, Toshio. (2007). Differences in neurogenic potential in floor plate cells along an anteroposterior location: Midbrain dopaminergic neurons originate from mesencephalic floor plate cells. Development, 134(17), 3213–3225. https://doi.org/10.1242/DEV.02879
Orii, Hidefumi; Kato, Kentaro; Agata, Kiyokazu; & Watanabe, Kenji. (1998). Molecular Cloning of Bone Morphogenetic Protein (BMP) Gene from the Planarian Dugesia japonica. Https://Doi.Org/10.2108/Zsj.15.871, 15(6), 871–877. https://doi.org/10.2108/ZSJ.15.871
Orii, Hidefumi; & Watanabe, Kenji. (2007). Bone morphogenetic protein is required for dorso-ventral patterning in the planarian Dugesia japonica. Development Growth and Differentiation, 49(4), 345–349. https://doi.org/10.1111/J.1440-169X.2007.00931.X,
Paisán-Ruíz, Coro; Jain, Shushant; Evans, E. Whitney; Gilks, William P.; Simón, Javier; Van Der Brug, Marcel; De Munain, Adolfo López; Aparicio, Silvia; Gil, Angel Martínez; Khan, Naheed; Johnson, Janel; Martinez, Javier Ruiz; Nicholl, David; Carrera, Itxaso Marti; Peňa, Amets Saénz; De Silva, Rohan; Lees, Andrew; Martí-Massó, José Félix; Pérez-Tur, Jordi; … Singleton, Andrew B. (2004). Cloning of the gene containing mutations that cause PARK8-linked Parkinson’s disease. Neuron, 44(4), 595–600. https://doi.org/10.1016/j.neuron.2004.10.023
Parkinson, James. (2002). An Essay on the Shaking Palsy. https://doi.org/10.1176/JNP.14.2.223
Pattyn, Alexandre; Morin, Xavier; Cremer, Harold; Goridis, Christo; & Brunet, Jean François. (1999). The homeobox gene Phox2b is essential for the development of autonomic neural crest derivatives. Nature, 399(6734), 366–370. https://doi.org/10.1038/20700
Pearse, A. G. (1969). The cytochemistry and ultrastructure of polypeptide hormone-producing cells of the APUD series and the embryologic, physiologic and pathologic implications of the concept. The Journal of Histochemistry and Cytochemistry : Official Journal of the Histochemistry Society, 17(5), 303–313. https://doi.org/10.1177/17.5.303
Pérez-Ferrer, Isabel; & Herrera, Eloísa. (2025). Novel insights into the mechanisms of growth cone dynamics during axon pathfinding. Current Opinion in Neurobiology, 93, 103073. https://doi.org/10.1016/J.CONB.2025.103073
Petersen, Christian P.; & Reddien, Peter W. (2008). Smed-βcatenin-1 is required for anteroposterior blastema polarity in planarian regeneration. Science, 319(5861), 327–330. https://doi.org/10.1126/science.1149943
Pla, Ramón; Borrell, Víctor; Flames, Nuria; & Marín, Oscar. (2006). Layer Acquisition by Cortical GABAergic Interneurons Is Independent of Reelin Signaling. Journal of Neuroscience, 26(26), 6924–6934. https://doi.org/10.1523/JNEUROSCI.0245-06.2006
Poliak, Sebastian; Norovich, Amy L.; Yamagata, Masahito; Sanes, Joshua R.; & Jessell, Thomas M. (2016). Muscle-type Identity of Proprioceptors Specified by Spatially Restricted Signals from Limb Mesenchyme. Cell, 164(3), 512–525. https://doi.org/10.1016/j.cell.2015.12.049
Polymeropoulos, Mihael H.; Lavedan, Christian; Leroy, Elisabeth; Ide, Susan E.; Dehejia, Anindya; Dutra, Amalia; Pike, Brian; Root, Holly; Rubenstein, Jeffrey; Boyer, Rebecca; Stenroos, Edward S.; Chandrasekharappa, Settara; Athanassiadou, Aglaia; Papapetropoulos, Theodore; Johnson, William G.; Lazzarini, Alice M.; Duvoisin, Roger C.; Di Iorio, Giuseppe; Golbe, Lawrence I.; & Nussbaum, Robert L. (1997). Mutation in the α-synuclein gene identified in families with Parkinson’s disease. Science, 276(5321), 2045–2047. https://doi.org/10.1126/SCIENCE.276.5321.2045
Pouchelon, Gabrielle; Gambino, Frédéric; Bellone, Camilla; Telley, Ludovic; Vitali, Ilaria; Lüscher, Christian; Holtmaat, Anthony; & Jabaudon, Denis. (2014). Modality-specific thalamocortical inputs instruct the identity of postsynaptic L4 neurons. Nature 2014 511:7510, 511(7510), 471–474. https://doi.org/10.1038/nature13390
Powell, Lynn M.; & Jarman, Andrew P. (2008). Context dependence of proneural bHLH proteins. Current Opinion in Genetics & Development, 18(5), 411. https://doi.org/10.1016/J.GDE.2008.07.012
Purves, Dale; Augustine, George J.; Fitzpatrick, David; Katz, Lawrence C.; LaMantia, Anthony-Samuel; McNamara, James O.; & Williams, S. Mark. (2001a). The Enteric Nervous System. https://www.ncbi.nlm.nih.gov/books/NBK11097/
Purves, Dale; Augustine, George J.; Fitzpatrick, David; Katz, Lawrence C.; LaMantia, Anthony-Samuel; McNamara, James O.; & Williams, S. Mark. (2001b). The Initial Formation of the Nervous System: Gastrulation and Neurulation. https://www.ncbi.nlm.nih.gov/books/NBK10993/
Qi, Cai; Luo, Li Da; Feng, Irena; & Ma, Shaojie. (2022). Molecular mechanisms of synaptogenesis. Frontiers in Synaptic Neuroscience, 14, 939793. https://doi.org/10.3389/FNSYN.2022.939793
Qualman, S. J.; Haupt, H. M.; Yang, P.; & Hamilton, S. R. (1984). Esophageal Lewy bodies associated with ganglion cell loss in achalasia. Similarity to Parkinson’s disease. Gastroenterology, 87(4), 848–856.
Quan, X. J.; & Hassan, B. A. (2005). From skin to nerve: flies, vertebrates and the first helix. Cellular and Molecular Life Sciences: CMLS, 62(18), 2036. https://doi.org/10.1007/S00018-005-5124-1
Que, Lin; Winterer, Jochen; & Földy, Csaba. (2019). Deep survey of GABAergic interneurons: Emerging insights from gene-isoform transcriptomics. Frontiers in Molecular Neuroscience, 12, 459023. https://doi.org/10.3389/fnmol.2019.00115
Raisman‐Vozari, Rita; Hirsch, Etienne; Javoy‐Agid, France; Vassort, Cecile; Savasta, Marc; Feuerstein, Claude; Thibault, Jean; & Agid, Yves. (1991). Quantitative Autoradiography of Tyrosine Hydroxylase Immunoreactivity in the Rat Brain. Journal of Neurochemistry, 57(4), 1212–1222. https://doi.org/10.1111/J.1471-4159.1991.TB08282.X
Raj, Bushra; Wagner, Daniel E.; McKenna, Aaron; Pandey, Shristi; Klein, Allon M.; Shendure, Jay; Gagnon, James A.; & Schier, Alexander F. (2018). Simultaneous single-cell profiling of lineages and cell types in the vertebrate brain. Nature Biotechnology 2018 36:5, 36(5), 442–450. https://doi.org/10.1038/nbt.4103
Rangel-Barajas, Claudia; Coronel, Israel; & Florán, Benjamín. (2015). Dopamine receptors and neurodegeneration. Aging and Disease, 6(5), 349–368. https://doi.org/10.14336/AD.2015.0330
Rasband, Matthew N. (2015). Glial Contributions to Neural Function and Disease. Molecular & Cellular Proteomics : MCP, 15(2), 355. https://doi.org/10.1074/MCP.R115.053744
Raz, Amelie A.; Wurtzel, Omri; & Reddien, Peter W. (2021). Planarian stem cells specify fate yet retain potency during the cell cycle. Cell Stem Cell, 28(7), 1307-1322.e5. https://doi.org/10.1016/j.stem.2021.03.021
Reddien, Peter W. (2018). The Cellular and Molecular Basis for Planarian Regeneration. Cell, 175(2), 327–345. https://doi.org/10.1016/j.cell.2018.09.021
Reddien, Peter W.; Bermange, Adam L.; Kicza, Adrienne M.; & Sánchez Alvarado, Alejandro. (2007). BMP signaling regulates the dorsal planarian midline and is needed for asymmetric regeneration. Development, 134(22), 4043–4051. https://doi.org/10.1242/dev.007138
Reddien, Peter W.; Oviedo, Néstor J.; Jennings, Joya R.; Jenkin, James C.; & Sánchez Alvarado, Alejandro. (2005). Developmental biology: SMEDWI-2 is a PIWI-like protein that regulates planarian stem cells. Science, 310(5752), 1327–1330. https://doi.org/10.1126/SCIENCE.1116110
Reddien, Peter W.; & Sánchez Alvarado, Alejandro. (2004). Fundamentals of planarian regeneration. Annual Review of Cell and Developmental Biology, 20, 725–757. https://doi.org/10.1146/annurev.cellbio.20.010403.095114
Richter, Sandra; Hartmann, Beate; & Reichert, Heinrich. (1998). The wingless gene is required for embryonic brain development in Drosophila. Development Genes and Evolution, 208(1), 37–45. https://doi.org/10.1007/S004270050151
Rink, Jochen C.; Vu, Hanh Thi Kim; & Alvarado, Alejandro Sánchez. (2011). The maintenance and regeneration of the planarian excretory system are regulated by EGFR signaling. Development, 138(17), 3769–3780. https://doi.org/10.1242/DEV.066852/-/DC1
Roberts-Galbraith, Rachel H.; & Newmark, Phillip A. (2015). On the organ trail: insights into organ regeneration in the planarian. Current Opinion in Genetics & Development, 32, 37–46. https://doi.org/10.1016/j.gde.2015.01.009
Rosenthal, Arnon. (1997). Specification and Survival of Dopaminergic Neurons in the Mammalian Midbrain. Advances in Pharmacology, 42(C), 908–911. https://doi.org/10.1016/S1054-3589(08)60894-7
Ross, Kelly G.; Currie, Ko W.; Pearson, Bret J.; & Zayas, Ricardo M. (2017). Nervous system development and regeneration in freshwater planarians. Wiley Interdisciplinary Reviews: Developmental Biology, 6(3). https://doi.org/10.1002/wdev.266
Rowlatt, Ursula. (1979). Intrauterine wound healing in a 20 week human fetus. Virchows Archiv A Pathological Anatomy and Histology, 381(3), 353–361. https://doi.org/10.1007/BF00432477
Sagner, Andreas; & Briscoe, James. (2019). Establishing neuronal diversity in the spinal cord: A time and a place. Development (Cambridge), 146(22). https://doi.org/10.1242/DEV.182154/223189
Sakai, Jill. (2020). How synaptic pruning shapes neural wiring during development and, possibly, in disease. Proceedings of the National Academy of Sciences, 117(28), 16096–16099. https://doi.org/10.1073/PNAS.2010281117
Salat, David; & Tolosa, Eduardo. (2013). Levodopa in the Treatment of Parkinson’s Disease: Current Status and New Developments. Journal of Parkinson’s Disease, 3(3), 255–269. https://doi.org/10.3233/JPD-130186
Saló, E.; & Baguñà, J. (1984). Regeneration and pattern formation in planarians. I. The pattern of mitosis in anterior and posterior regeneration in Dugesia (G) tigrina, and a new proposal for blastema formation. Journal of Embryology and Experimental Morphology, 83, 63–80.
Sánchez-Pernaute, Rosario; Studer, Lorenz; Bankiewicz, Krys S.; Major, Eugene O.; & McKay, Ronald D. G. (2001). In vitro generation and transplantation of precursor-derived human dopamine neurons. Journal of Neuroscience Research, 65(4), 284–288. https://doi.org/10.1002/JNR.1152
Sasai, Y. (2001). Roles of Sox factors in neural determination: conserved signaling in evolution? The International Journal of Developmental Biology, 45(1), 321–326.
Satapathy, Sandeep; & Wilson, Mark R. (2022). Roles of constitutively secreted extracellular chaperones in neuronal cell repair and regeneration. Neural Regeneration Research, 18(4), 769. https://doi.org/10.4103/1673-5374.353483
Sawamoto, Nobukatsu; Doi, Daisuke; Nakanishi, Etsuro; Sawamura, Masanori; Kikuchi, Takayuki; Yamakado, Hodaka; Taruno, Yosuke; Shima, Atsushi; Fushimi, Yasutaka; Okada, Tomohisa; Kikuchi, Tetsuhiro; Morizane, Asuka; Hiramatsu, Satoe; Anazawa, Takayuki; Shindo, Takero; Ueno, Kentaro; Morita, Satoshi; Arakawa, Yoshiki; Nakamoto, Yuji; … Takahashi, Jun. (2025). Phase I/II trial of iPS-cell-derived dopaminergic cells for Parkinson’s disease. Nature 2025 641:8064, 641(8064), 971–977. https://doi.org/10.1038/s41586-025-08700-0
Schier, Alexander F.; & Talbot, William S. (2005). Molecular genetics of axis formation in zebrafish. Annual Review of Genetics, 39(Volume 39, 2005), 561–613. https://doi.org/10.1146/annurev.genet.37.110801.143752
Scimone, M. Lucila; Srivastava, Mansi; Bell, George W.; & Reddien, Peter W. (2011). A regulatory program for excretory system regeneration in planarians. Development, 138(20), 4387–4398. https://doi.org/10.1242/DEV.068098
Scimone, M. Lucila; Wurtzel, Omri; Malecek, Kathryn; Fincher, Christopher T.; Oderberg, Isaac M.; Kravarik, Kellie M.; & Reddien, Peter W. (2018). foxF-1 Controls Specification of Non-body Wall Muscle and Phagocytic Cells in Planarians. Current Biology, 28(23), 3787-3801.e6. https://doi.org/10.1016/J.CUB.2018.10.030
Sethi, Kapil D. (2010). The impact of levodopa on quality of life in patients with Parkinson disease. Neurologist, 16(2), 76–83. https://doi.org/10.1097/NRL.0B013E3181BE6D15
Sharma, Aditi; Shah, Om Prakash; Sharma, Lalit; Gulati, Monica; Behl, Tapan; Khalid, Asaad; Mohan, Syam; Najmi, Asim; & Zoghebi, Khalid. (2024). Molecular Chaperones as Therapeutic Target: Hallmark of Neurodegenerative Disorders. Molecular Neurobiology, 61(7), 4750–4767. https://doi.org/10.1007/S12035-023-03846-2
Shusharina, Natalia; Yukhnenko, Denis; Botman, Stepan; Sapunov, Viktor; Savinov, Vladimir; Kamyshov, Gleb; Sayapin, Dmitry; & Voznyuk, Igor. (2023). Modern Methods of Diagnostics and Treatment of Neurodegenerative Diseases and Depression. Diagnostics, 13(3), 573. https://doi.org/10.3390/DIAGNOSTICS13030573
Sidransky, E.; Nalls, M. A.; Aasly, J. O.; Aharon-Peretz, J.; Annesi, G.; Barbosa, E. R.; Bar-Shira, A.; Berg, D.; Bras, J.; Brice, A.; Chen, C. M.; Clark, L. N.; Condroyer, C.; De Marco, E. V.; Dürr, A.; Eblan, M. J.; Fahn, S.; Farrer, M. J.; Fung, H. C.; … Ziegler, S. G. (2009). Multicenter Analysis of Glucocerebrosidase Mutations in Parkinson’s Disease. New England Journal of Medicine, 361(17), 1651–1661. https://doi.org/10.1056/NEJMOA0901281
Sidransky, Ellen; & Lopez, Grisel. (2012). The link between the GBA gene and parkinsonism. The Lancet Neurology, 11(11), 986–998. https://doi.org/10.1016/S1474-4422(12)70190-4
Siehr, Meagan S.; Koo, Pamela K.; Sherlekar, Amrita L.; Bian, Xuelin; Bunkers, Meredith R.; Miller, Renee M.; Portman, Douglas S.; & Lints, Robyn. (2011). Multiple doublesex-related genes specify critical cell fates in a C. elegans male neural circuit. PLoS ONE, 6(10). https://doi.org/10.1371/JOURNAL.PONE.0026811
Siletti, Kimberly; Hodge, Rebecca; Mossi Albiach, Alejandro; Lee, Ka Wai; Ding, Song Lin; Hu, Lijuan; Lönnerberg, Peter; Bakken, Trygve; Casper, Tamara; Clark, Michael; Dee, Nick; Gloe, Jessica; Hirschstein, Daniel; Shapovalova, Nadiya V.; Dirk Keene, C.; Nyhus, Julie; Tung, Herman; Yanny, Anna Marie; Arenas, Ernest; … Linnarsson, Sten. (2023). Transcriptomic diversity of cell types across the adult human brain. Science, 382(6667). https://doi.org/10.1126/SCIENCE.ADD7046
Silvin, Aymeric; & Ginhoux, Florent. (2018). Microglia heterogeneity along a spatio–temporal axis: More questions than answers. Glia, 66(10), 2045–2057. https://doi.org/10.1002/GLIA.23458
Sinigaglia, Chiara; Busengdal, Henriette; Leclère, Lucas; Technau, Ulrich; & Rentzsch, Fabian. (2013). The Bilaterian Head Patterning Gene six3/6 Controls Aboral Domain Development in a Cnidarian. PLOS Biology, 11(2), e1001488. https://doi.org/10.1371/JOURNAL.PBIO.1001488
Skeath, James B.; Zhang, Yu; Holmgren, Robert; Carroll, Sean B.; & Doe, Chris Q. (1995). Specification of neuroblast identity in the drosophila embryonic central nervous system by gooseberry-distal. Nature, 376(6539), 427–430. https://doi.org/10.1038/376427a0
Smidt, Marten P.; Asbreuk, Ceriel H. J.; Cox, Joke J.; Chen, Haixu; Johnson, Randy L.; & Burbach, J. Peter H. (2000). A second independent pathway for development of mesencephalic dopaminergic neurons requires Lmx1b. Nature Neuroscience, 3(4), 337–341. https://doi.org/10.1038/73902
Smith, Heather L.; Li, Wenwen; & Cheetham, Michael E. (2015). Molecular chaperones and neuronal proteostasis. Seminars in Cell & Developmental Biology, 40, 142. https://doi.org/10.1016/J.SEMCDB.2015.03.003
Smith, William C.; & Harland, Richard M. (1992). Expression cloning of noggin, a new dorsalizing factor localized to the Spemann organizer in Xenopus embryos. Cell, 70(5), 829–840. https://doi.org/10.1016/0092-8674(92)90316-5
Sonntag, Kai-Christian; Pruszak, Jan; Yoshizaki, Takahito; van Arensbergen, Joris; Sanchez-Pernaute, Rosario; & Isacson, Ole. (2007). Enhanced Yield of Neuroepithelial Precursors and Midbrain-Like Dopaminergic Neurons from Human Embryonic Stem Cells Using the Bone Morphogenic Protein Antagonist Noggin. Stem Cells, 25(2), 411–418. https://doi.org/10.1634/STEMCELLS.2006-0380
Sousa, Sara C.; & Sousa, Mónica M. (2021). The cytoskeleton as a modulator of tension driven axon elongation. Developmental Neurobiology, 81(3), 300–309. https://doi.org/10.1002/DNEU.22747
Sreedharan, Jemeen; Blair, Ian P.; Tripathi, Vineeta B.; Hu, Xun; Vance, Caroline; Rogelj, Boris; Ackerley, Steven; Durnall, Jennifer C.; Williams, Kelly L.; Buratti, Emanuele; Baralle, Francisco; De Belleroche, Jacqueline; Mitchell, J. Douglas; Leigh, P. Nigel; Al-Chalabi, Ammar; Miller, Christopher C.; Nicholson, Garth; & Shaw, Christopher E. (2008). TDP-43 mutations in familial and sporadic amyotrophic lateral sclerosis. Science (New York, N.Y.), 319(5870), 1668–1672. https://doi.org/10.1126/SCIENCE.1154584
Srinivasan, E.; Chandrasekhar, G.; Chandrasekar, P.; Anbarasu, K.; Vickram, A. S.; Karunakaran, Rohini; Rajasekaran, R.; & Srikumar, P. S. (2021). Alpha-Synuclein Aggregation in Parkinson’s Disease. Frontiers in Medicine, 8, 736978. https://doi.org/10.3389/FMED.2021.736978
Storer, Mekayla A.; & Miller, Freda D. (2020). Cellular and molecular mechanisms that regulate mammalian digit tip regeneration: Mechanisms of Digit Tip Regeneration. Open Biology, 10(9). https://doi.org/10.1098/RSOB.200194/
Stratoulias, Vassilis; Venero, Jose Luis; Tremblay, Marie‐Ève; & Joseph, Bertrand. (2019). Microglial subtypes: diversity within the microglial community. The EMBO Journal, 38(17). https://doi.org/10.15252/EMBJ.2019101997
Sugino, Ken; Clark, Erin; Schulmann, Anton; Shima, Yasuyuki; Wang, Lihua; Hunt, David L.; Hooks, Bryan M.; Nkner, Dimitri Tra¨; Chandrashekar, Jayaram; Picard, Serge; Lemire, Andrew L.; Spruston, Nelson; Hantman, Adam W.; & Nelson, Sacha B. (2019). Mapping the transcriptional diversity of genetically and anatomically defined cell populations in the mouse brain. ELife, 8. https://doi.org/10.7554/ELIFE.38619
Suk, Terry R.; & Rousseaux, Maxime W. C. (2020). The role of TDP-43 mislocalization in amyotrophic lateral sclerosis. Molecular Neurodegeneration, 15(1), 1–16. https://doi.org/10.1186/s13024-020-00397-1
Sulston, J.; Dew, M.; & Brenner, S. (1975). Dopaminergic neurons in the nematode Caenorhabditis elegans. Journal of Comparative Neurology, 163(2), 215–226. https://doi.org/10.1002/CNE.901630207
Sulston, J. E. & Horvitz, H. R. (1977). Post-embryonic cell lineages of the nematode, Caenorhabditis elegans. Developmental Biology, 56(1), 110–156. https://doi.org/10.1016/0012-1606(77)90158-0
Sulston, J. E.; Schierenberg, E.; White, J. G.; & Thomson, J. N. (1983). The embryonic cell lineage of the nematode Caenorhabditis elegans. Developmental Biology, 100(1), 64–119. https://doi.org/10.1016/0012-1606(83)90201-4
Sweeney, Patrick; Park, Hyunsun; Baumann, Marc; Dunlop, John; Frydman, Judith; Kopito, Ron; McCampbell, Alexander; Leblanc, Gabrielle; Venkateswaran, Anjli; Nurmi, Antti; & Hodgson, Robert. (2017). Protein misfolding in neurodegenerative diseases: implications and strategies. Translational Neurodegeneration, 6(1), 6. https://doi.org/10.1186/S40035-017-0077-5
Tabar, V.;, Sarva, H.; Lozano, A. M.; Fasano, A.; Kalia, S. K.; Yu, K. K. H.; Brennan, C.; Ma, Y.; Peng, S.; Eidelberg, D.; Tomishima, M.; Irion, S.; Stemple, W.; Abid, N.; Lampron, A.; Studer, L.; & Henchcliffe, C. (2025). Phase I trial of hES cell-derived dopaminergic neurons for Parkinson’s disease. Nature, 641(8064), 978–983. https://doi.org/10.1038/s41586-025-08845-y
Takahashi, Kazutoshi; Tanabe, Koji; Ohnuki, Mari; Narita, Megumi; Ichisaka, Tomoko; Tomoda, Kiichiro; & Yamanaka, Shinya. (2007). Induction of Pluripotent Stem Cells from Adult Human Fibroblasts by Defined Factors. Cell, 131(5), 861–872. https://doi.org/10.1016/j.cell.2007.11.019
Takeda, Hiroyuki; Nishimura, Kaneyasu; & Agata, Kiyokazu. (2009). Planarians maintain a constant ratio of different cell types during changes in body size by using the stem cell system. Zoological Science, 26(12), 805–813. https://doi.org/10.2108/ZSJ.26.805
Takeo, Makoto; Lee, Wendy; & Ito, Mayumi. (2015). Wound Healing and Skin Regeneration. Cold Spring Harbor Perspectives in Medicine, 5(1), a023267. https://doi.org/10.1101/CSHPERSPECT.A023267
Tan, Yun Long; Yuan, Yi; & Tian, Li. (2020). Microglial regional heterogeneity and its role in the brain. Molecular Psychiatry, 25(2), 351–367. https://doi.org/10.1038/s41380-019-0609-8
Tanaka, Elly M.; & Reddien, Peter W. (2011). The cellular basis for animal regeneration. Developmental Cell, 21(1), 172. https://doi.org/10.1016/J.DEVCEL.2011.06.016
Taraviras, Stavros; Marcos-Gutierrez, Camelia V.; Durbec, Pascale; Jani, Harsha; Grigoriou, Maria; Sukumaran, Madhu; Wang, Li Chong; Hynes, Mary; Raisman, Geoffrey; & Pachnis, Vassilis. (1999). Signalling by the RET receptor tyrosine kinase and its role in the development of the mammalian enteric nervous system. Development, 126(12), 2785–2797. https://doi.org/10.1242/DEV.126.12.2785
Taverna, Elena; Götz, Magdalena; & Huttner, Wieland B. (2014). The cell biology of neurogenesis: toward an understanding of the development and evolution of the neocortex. Annual Review of Cell and Developmental Biology, 30(Volume 30, 2014), 465–502. https://doi.org/10.1146/annurev-cellbio-101011-155801
Telley, L.; Agirman, G.; Prados, J.; Amberg, N.; Fièvre, S.; Oberst, P.; Bartolini, G.; Vitali, I.; Cadilhac, C.; Hippenmeyer, S.; Nguyen, L.; Dayer, A.; & Jabaudon, D. (2019). Temporal patterning of apical progenitors and their daughter neurons in the developing neocortex. Science, 364(6440). https://doi.org/10.1126/science.aav2522
Thomson, James A. (1998). Embryonic stem cell lines derived from human blastocysts. Science, 282(5391), 1145–1147. https://doi.org/10.1126/SCIENCE.282.5391.1145,
Tolosa, Eduardo S.; Martin, William E.; Cohen, Harold P.; & Jacobson, Ronald L. (1975). Patterns of clinical response and plasma dopa levels in parkinson’s disease. Neurology, 25(2), 177–183. https://doi.org/10.1212/WNL.25.2.177
Trevisan, L.; Gaudio, A.; Monfrini, E.; Avanzino, L.; Di Fonzo, A.; & Mandich, P. (2024). Genetics in Parkinson’s disease, state-of-the-art and future perspectives. British Medical Bulletin, 149(1), 60. https://doi.org/10.1093/BMB/LDAD035
Tu, Kimberly C.; Cheng, Li Chun; Vu, Hanh T. K.; Lange, Jeffrey J.; McKinney, Sean A.; Seidel, Chris W.; & Sánchez Alvarado, Alejandro. (2015). Egr-5 is a post-mitotic regulator of planarian epidermal differentiation. ELife, 4(OCTOBER2015), e10501. https://doi.org/10.7554/ELIFE.10501
Ulvklo, Carina; MacDonald, Ryan; Bivik, Caroline; Baumgardt, Magnus; Karlsson, Daniel; & Thor, Stefan. (2012). Control of neuronal cell fate and number by integration of distinct daughter cell proliferation modes with temporal progression. Development, 139(4), 678–689. https://doi.org/10.1242/DEV.074500
Umesono, Yoshihiko; & Agata, Kiyokazu. (2009). Evolution and regeneration of the planarian central nervous system. Development, Growth & Differentiation, 51(3), 185–195. https://doi.org/10.1111/J.1440-169X.2009.01099.X
Ungerstedt, U.; Ljungberg, T.; & Steg, G. (1974). Behavioral, physiological, and neurochemical changes after 6-hydroxydopamine-induced degeneration of the nigro-striatal dopamine neurons. Advances in Neurology, 5, 421–426.
Valadez-Barba, Valeria; Cota-Coronado, A.; Hernández-Pérez, O. R.; Lugo-Fabres, Pavel H.; Padilla-Camberos, Eduardo; Díaz, Néstor Fabián; & Díaz-Martínez, N. Emmanuel. (2020). iPSC for modeling neurodegenerative disorders. Regenerative Therapy, 15, 332–339. https://doi.org/10.1016/J.RETH.2020.11.006
van Wolfswinkel, Josien C.; Wagner, Daniel E.; & Reddien, Peter W. (2014). Single-Cell Analysis Reveals Functionally Distinct Classes within the Planarian Stem Cell Compartment. Cell Stem Cell, 15(3), 326–339. https://doi.org/10.1016/J.STEM.2014.06.007
Varadarajan, Supraja G.; Hunyara, John L.; Hamilton, Natalie R.; Kolodkin, Alex L.; & Huberman, Andrew D. (2022). Central nervous system regeneration. Cell, 185(1), 77–94. https://doi.org/10.1016/J.CELL.2021.10.029
Wagner, Daniel E.; Wang, Irving E.; & Reddien, Peter W. (2011). Clonogenic Neoblasts Are Pluripotent Adult Stem Cells That Underlie Planarian Regeneration. Science, 332(6031), 811–816. https://doi.org/10.1126/science.1203983
Wahbe, F.; Hagege, J.; Loreau, N.; & Ardaillou, R. (1982). Endogenous dopamine synthesis and DOPA-decarboxylase activity in rat renal cortex. Molecular and Cellular Endocrinology, 27(1), 45–54. https://doi.org/10.1016/0303-7207(82)90061-2
Wakabayashi, K.; Takahashi, H.; Takeda, S.; Ohama, E.; & Ikuta, F. (1988). Parkinson’s disease: the presence of Lewy bodies in Auerbach’s and Meissner’s plexuses. Acta Neuropathologica, 76(3), 217–221. https://doi.org/10.1007/BF00687767
Walkers, R. J.; & Holden-Dye, Lindy. (1989). Commentary on the evolution of transmitters, receptors and ion channels in invertebrates. Comparative Biochemistry and Physiology — Part A: Physiology, 93(1), 25–39. https://doi.org/10.1016/0300-9629(89)90188-6
Wang, Irving E.; Lapan, Sylvain W.; Scimone, M. Lucila; Clandinin, Thomas R.; & Reddien, Peter W. (2016). Hedgehog signaling regulates gene expression in planarian glia. ELife, 5(September2016). https://doi.org/10.7554/ELIFE.16996
Watanabe, Hiroshi; Fujisawa, Toshitaka; & Holstein, Thomas W. (2009). Cnidarians and the evolutionary origin of the nervous system. Development, Growth & Differentiation, 51(3), 167–183. https://doi.org/10.1111/J.1440-169X.2009.01103.X
Watanabe, Hiroshi; Hoang, Van Thanh; Mättner, Robert; & Holstein, Thomas W. (2009). Immortality and the base of multicellular life: Lessons from cnidarian stem cells. Seminars in Cell & Developmental Biology, 20(9), 1114–1125. https://doi.org/10.1016/J.SEMCDB.2009.09.008
Watanabe, Yuli; Stanchina, Laure; Lecerf, Laure; Gacem, Nadjet; Conidi, Andrea; Baral, Viviane; Pingault, Veronique; Huylebroeck, Danny; & Bondurand, Nadege. (2017). Differentiation of Mouse Enteric Nervous System Progenitor Cells Is Controlled by Endothelin 3 and Requires Regulation of Ednrb by SOX10 and ZEB2. Gastroenterology, 152(5), 1139-1150.e4. https://doi.org/10.1053/J.GASTRO.2016.12.034
Wei, Yonglong; Wang, Yunguan G.; Jia, Yuemeng; Li, Lin; Yoon, Jung; Zhang, Shuyuan; Wang, Zixi; Zhang, Yu; Zhu, Min; Sharma, Tripti; Lin, Yu Hsuan; Hsieh, Meng Hsiung;, Albrecht, Jeffrey H.; Le, Phuong T.; Rosen, Clifford J.; Wang, Tao; & Zhu, Hao. (2021). Liver homeostasis is maintained by midlobular zone 2 hepatocytes. Science, 371(6532). https://doi.org/10.1126/science.abb1625
Wenemoser, Danielle; & Reddien, Peter W. (2010). Planarian regeneration involves distinct stem cell responses to wounds and tissue absence. Developmental Biology, 344(2), 979–991. https://doi.org/10.1016/j.ydbio.2010.06.017
Westergard, Thomas; & Rothstein, Jeffrey D. (2020). Astrocyte Diversity: Current Insights and Future Directions. Neurochemical Research, 45(6), 1298–1305. https://doi.org/10.1007/s11064-020-02959-7
White, John; Southgate, E.; Thomson, J.; & Brenner, S. (1986). The structure of the nervous system of the nematode Caenorhabditis elegans. Philosophical Transactions of the Royal Society of London. B, Biological Sciences, 314(1165), 1–340. https://doi.org/10.1098/rstb.1986.0056
Wictorin, K.; Brundin, P.; Sauer, H.; Lindvall, O.; & Björklund, A. (1992). Long distance directed axonal growth from human dopaminergic mesencephalic neuroblasts implanted along the nigrostriatal pathway in 6‐hydroxydopamine lesioned adult rats. Journal of Comparative Neurology, 323(4), 475–494. https://doi.org/10.1002/CNE.903230403
Widodo, Wahyu; Aprilya, Dina; & Satria, Oryza. (2025). Regenerative Medicine: A New Horizon in Peripheral Nerve Injury and Repair. Orthopedic Reviews, 17, 2025. https://doi.org/10.52965/001C.133572
Wolterhoff, Neele; & Hiesinger, P. Robin. (2024). Synaptic promiscuity in brain development. Current Biology, 34(3), R102–R116. https://doi.org/10.1016/J.CUB.2023.12.037
Xiong, Man; Tao, Yezheng; Gao, Qinqin; Feng, Ban; Yan, Wei; Zhou, Yingying; Kotsonis, Thomas A.; Yuan, Tingli; You, Zhiwen; Wu, Ziyan; Xi, Jiajie; Haberman, Alexander; Graham, Julia; Block, Jasper; Zhou, Wenhao; Chen, Yuejun; & Zhang, Su Chun. (2021). Human Stem Cell-Derived Neurons Repair Circuits and Restore Neural Function. Cell Stem Cell, 28(1), 112-126.e6. https://doi.org/10.1016/j.stem.2020.08.014
Yang, Dali; Zhang, Zhi-Jian; Oldenburg, Michael; Ayala, Melvin; & Zhang, Su-Chun. (2008). Human Embryonic Stem Cell-Derived Dopaminergic Neurons Reverse Functional Deficit in Parkinsonian Rats. Stem Cells, 26(1), 55–63. https://doi.org/10.1634/STEMCELLS.2007-0494
Yntema, Chester L.; & Hammond, Warner S. (1954). The origin of intrinsic ganglia of trunk viscera from vagal neural crest in the chick embryo. Journal of Comparative Neurology, 101(2), 515–541. https://doi.org/10.1002/CNE.901010212
Yntema, Chester L.; & Hammond, Warner S. (1955). Experiments on the origin and development of the sacral autonomic nerves in the chick embryo. Journal of Experimental Zoology, 129(2), 375–413. https://doi.org/10.1002/JEZ.1401290210
Zetterström, Rolf H.; Williams, Reg; Perlmann, Thomas; & Olson, Lars. (1996). Cellular expression of the immediate early transcription factors Nurr1 and NGFI-B suggests a gene regulatory role in several brain regions including the nigrostriatal dopamine system. Molecular Brain Research, 41(1–2), 111–120. https://doi.org/10.1016/0169-328X(96)00074-5
Zhang, Su Chun; Wernig, Marius; Duncan, Ian D.; Brüstle, Oliver; & Thomson, James A. (2001). In vitro differentiation of transplantable neural precursors from human embryonic stem cells. Nature Biotechnology, 19(12), 1129–1133. https://doi.org/10.1038/NBT1201-1129
Zhu, Shu Jun; & Pearson, Bret J. (2016). (Neo)blast from the past: new insights into planarian stem cell lineages. Current Opinion in Genetics and Development, 40, 74–80. https://doi.org/10.1016/j.gde.2016.06.007
Zimprich, Alexander; Biskup, Saskia; Leitner, Petra; Lichtner, Peter; Farrer, Matthew; Lincoln, Sarah; Kachergus, Jennifer; Hulihan, Mary; Uitti, Ryan J.; Calne, Donald B.; Stoessl, A. Jon; Pfeiffer, Ronald F.; Patenge, Nadja; Carbajal, Iria Carballo; Vieregge, Peter; Asmus, Friedrich; Müller-Myhsok, Bertram; Dickson, Dennis W.; Meitinger, Thomas; … Gasser, Thomas. (2004). Mutations in LRRK2 cause autosomal-dominant parkinsonism with pleomorphic pathology. Neuron, 44(4), 601–607. https://doi.org/10.1016/j.neuron.2004.11.005

